| Record Information |
|---|
| Version | 1.0 |
|---|
| Created at | 2020-04-17 18:49:39 UTC |
|---|
| Updated at | 2020-12-07 19:11:15 UTC |
|---|
| CannabisDB ID | CDB004911 |
|---|
| Secondary Accession Numbers | Not Available |
|---|
| Cannabis Compound Identification |
|---|
| Common Name | Guanosine monophosphate |
|---|
| Description | Guanosine monophosphate, also known as guanylic acid or 5'-GMP, belongs to the class of organic compounds known as purine ribonucleoside monophosphates. These are nucleotides consisting of a purine base linked to a ribose to which one monophosphate group is attached. Guanosine monophosphate is an extremely weak basic (essentially neutral) compound (based on its pKa). Guanosine monophosphate exists in all living species, ranging from bacteria to humans. Within humans, guanosine monophosphate participates in a number of enzymatic reactions. In particular, guanosine triphosphate and guanosine monophosphate can be biosynthesized from diguanosine tetraphosphate; which is catalyzed by the enzyme bis(5'-nucleosyl)-tetraphosphatase [asymmetrical]. In addition, guanosine monophosphate can be biosynthesized from guanosine diphosphate; which is mediated by the enzyme ectonucleoside triphosphate diphosphohydrolase 5. In humans, guanosine monophosphate is involved in purine metabolism. A purine ribonucleoside 5'-monophosphate having guanine as the nucleobase. Outside of the human body, Guanosine monophosphate has been detected, but not quantified in, several different foods, such as rosemaries, common chokecherries, pigeon pea, rices, and horseradish tree. This could make guanosine monophosphate a potential biomarker for the consumption of these foods. Guanosine monophosphate is expected to be in Cannabis as all living plants are known to produce and metabolize it. |
|---|
| Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| 5'-GMP | ChEBI | | GMP | ChEBI | | Guanosine 5'-phosphate | ChEBI | | Guanosine-5'-monophosphate | ChEBI | | Guanylic acid | ChEBI | | pG | ChEBI | | Guanosine 5'-monophosphate | Kegg | | Guanosine 5'-phosphoric acid | Generator | | Guanosine-5'-monophosphoric acid | Generator | | Guanylate | Generator | | Guanosine 5'-monophosphoric acid | Generator | | Guanosine monophosphoric acid | Generator | | e 626 | HMDB | | Guanidine monophosphate | HMDB | | Guanosine 5'-phosphorate | HMDB | | Guanosine-5'-phosphate | HMDB | | Guanosine-phosphate | HMDB | | 5' Guanylic acid | HMDB | | Acid, 5'-guanylic | HMDB | | Acid, guanylic | HMDB | | Guanosine 5' monophosphate | HMDB | | 5'-monoPhosphate, guanosine | HMDB | | monoPhosphate, guanosine | HMDB | | 5'-Guanylic acid | HMDB |
|
|---|
| Chemical Formula | C10H14N5O8P |
|---|
| Average Molecular Weight | 363.22 |
|---|
| Monoisotopic Molecular Weight | 363.058 |
|---|
| IUPAC Name | {[(2R,3S,4R,5R)-5-(2-amino-6-oxo-6,9-dihydro-1H-purin-9-yl)-3,4-dihydroxyoxolan-2-yl]methoxy}phosphonic acid |
|---|
| Traditional Name | guanylate |
|---|
| CAS Registry Number | 85-32-5 |
|---|
| SMILES | NC1=NC2=C(N=CN2[C@@H]2O[C@H](COP(O)(O)=O)[C@@H](O)[C@H]2O)C(=O)N1 |
|---|
| InChI Identifier | InChI=1S/C10H14N5O8P/c11-10-13-7-4(8(18)14-10)12-2-15(7)9-6(17)5(16)3(23-9)1-22-24(19,20)21/h2-3,5-6,9,16-17H,1H2,(H2,19,20,21)(H3,11,13,14,18)/t3-,5-,6-,9-/m1/s1 |
|---|
| InChI Key | RQFCJASXJCIDSX-UUOKFMHZSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | Belongs to the class of organic compounds known as purine ribonucleoside monophosphates. These are nucleotides consisting of a purine base linked to a ribose to which one monophosphate group is attached. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Nucleosides, nucleotides, and analogues |
|---|
| Class | Purine nucleotides |
|---|
| Sub Class | Purine ribonucleotides |
|---|
| Direct Parent | Purine ribonucleoside monophosphates |
|---|
| Alternative Parents | |
|---|
| Substituents | - Purine ribonucleoside monophosphate
- Pentose phosphate
- Pentose-5-phosphate
- Glycosyl compound
- N-glycosyl compound
- 6-oxopurine
- Hypoxanthine
- Monosaccharide phosphate
- Pentose monosaccharide
- Imidazopyrimidine
- Purine
- Aminopyrimidine
- Monoalkyl phosphate
- Pyrimidone
- Alkyl phosphate
- Pyrimidine
- Monosaccharide
- N-substituted imidazole
- Organic phosphoric acid derivative
- Phosphoric acid ester
- Tetrahydrofuran
- Azole
- Imidazole
- Heteroaromatic compound
- Vinylogous amide
- 1,2-diol
- Secondary alcohol
- Oxacycle
- Azacycle
- Organoheterocyclic compound
- Primary amine
- Organic nitrogen compound
- Amine
- Hydrocarbon derivative
- Organic oxide
- Organopnictogen compound
- Organic oxygen compound
- Alcohol
- Organonitrogen compound
- Organooxygen compound
- Aromatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aromatic heteropolycyclic compounds |
|---|
| External Descriptors | |
|---|
| Ontology |
|---|
|
| Disposition | Source: Biological location: |
|---|
| Role | Industrial application: Biological role: |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Experimental Properties | | Property | Value | Reference |
|---|
| Melting Point | Not Available | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | 369 mg/mL | Not Available | | logP | Not Available | Not Available |
|
|---|
| Predicted Properties | [] |
|---|
| Spectra |
|---|
| EI-MS/GC-MS | | Type | Description | Splash Key | View |
|---|
| GC-MS | Guanosine monophosphate, 6 TMS, GC-MS Spectrum | splash10-014i-1942000000-27fc2e135f86071ba4d7 | Spectrum | | GC-MS | Guanosine monophosphate, non-derivatized, GC-MS Spectrum | splash10-014i-1942000000-27fc2e135f86071ba4d7 | Spectrum | | GC-MS | Guanosine monophosphate, non-derivatized, GC-MS Spectrum | splash10-014i-0941000000-245c5258141d8d2a9fa5 | Spectrum | | Predicted GC-MS | Guanosine monophosphate, non-derivatized, Predicted GC-MS Spectrum - 70eV, Positive | splash10-0002-9722000000-9155df338c7516c297b6 | Spectrum | | Predicted GC-MS | Guanosine monophosphate, 2 TMS, Predicted GC-MS Spectrum - 70eV, Positive | splash10-01ot-9522200000-3e2591b21841c753ec13 | Spectrum | | Predicted GC-MS | Guanosine monophosphate, non-derivatized, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum |
|
|---|
| MS/MS | | Type | Description | Splash Key | View |
|---|
| MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 10V, Positive (Annotated) | splash10-0udi-0901000000-fbce35f7dfa73ab5fc11 | 2012-07-24 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 25V, Positive (Annotated) | splash10-0udi-0900000000-764722cc1b8cc2aaa480 | 2012-07-24 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 40V, Positive (Annotated) | splash10-0udi-0900000000-c268cdce8dfd6c39fa3c | 2012-07-24 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF (UPLC Q-Tof Premier, Waters) , Positive | splash10-03di-0409000000-309e438ad8b6162df850 | 2012-08-31 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF (UPLC Q-Tof Premier, Waters) 30V, Positive | splash10-0udi-0904000000-cf72d0459316801b014e | 2012-08-31 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF (UPLC Q-Tof Premier, Waters) , Positive | splash10-0udi-0900000000-abc7bfe7620db1c34775 | 2012-08-31 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF (UPLC Q-Tof Premier, Waters) , Negative | splash10-01t9-9114000000-9a1071a2e17ea3ca268c | 2012-08-31 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF (UPLC Q-Tof Premier, Waters) , Negative | splash10-01t9-9113000000-05a8989070e3b89138f1 | 2012-08-31 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , negative | splash10-01t9-9114000000-9a1071a2e17ea3ca268c | 2017-09-14 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , negative | splash10-01t9-9113000000-05a8989070e3b89138f1 | 2017-09-14 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , positive | splash10-03di-0409000000-309e438ad8b6162df850 | 2017-09-14 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , positive | splash10-0udi-0904000000-cf72d0459316801b014e | 2017-09-14 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , positive | splash10-0udi-0900000000-abc7bfe7620db1c34775 | 2017-09-14 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - 40V, Negative | splash10-004i-9000000000-c340c748805ec82deb0d | 2021-09-20 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - 10V, Negative | splash10-03di-3009000000-5d594db970a4fea1099e | 2021-09-20 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - 20V, Negative | splash10-004i-9011000000-8335076473133728e58f | 2021-09-20 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - 30V, Positive | splash10-0udi-0904000000-cf72d0459316801b014e | 2021-09-20 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - 20V, Positive | splash10-0udi-0900000000-1f479e6ff94baf0265bc | 2021-09-20 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - 40V, Positive | splash10-0udi-1900000000-1dd27aac9ee9a0882191 | 2021-09-20 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0udi-0913000000-05cd931f142e4b624c8b | 2016-09-12 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0udi-0900000000-318d089a94a5bcfd20f2 | 2016-09-12 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0udi-0900000000-e616291ba80c4abf405c | 2016-09-12 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0imi-7709000000-6c811759b6f078f642a2 | 2016-09-12 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0fb9-9800000000-59c4ae8bf5bfa0d23da1 | 2016-09-12 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-004i-9100000000-73f60c52b80206170819 | 2016-09-12 | View Spectrum |
|
|---|
| NMR | | Type | Description | | View |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 500 MHz, H2O, experimental) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 100 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 100 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 200 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 200 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 300 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 300 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 400 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 500 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 500 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 600 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 600 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 700 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 700 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 800 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 800 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 900 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 900 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 1000 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, D2O, predicted) | | Spectrum | | 2D NMR | [1H, 13C]-HSQC NMR Spectrum (2D, 600 MHz, H2O, experimental) | | Spectrum |
|
|---|
| Pathways |
|---|
| Pathways | | Name | SMPDB/Pathwhiz | KEGG | | Transcription/Translation | Not Available | Not Available | | Purine Metabolism |  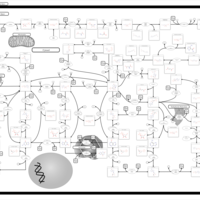 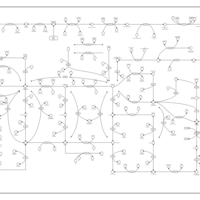 |  | | Glutamate Metabolism | 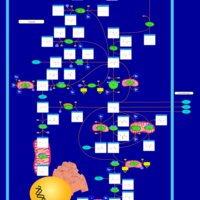 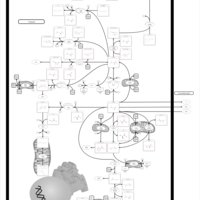 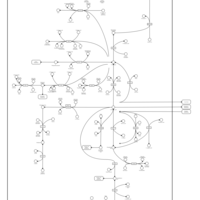 |  | | Adenosine Deaminase Deficiency |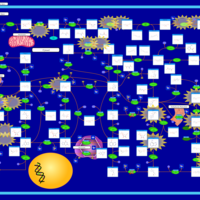  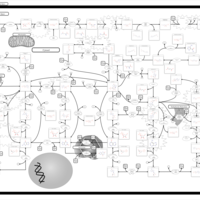 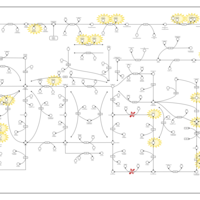 | Not Available | | Adenylosuccinate Lyase Deficiency | 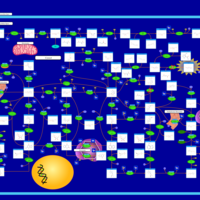 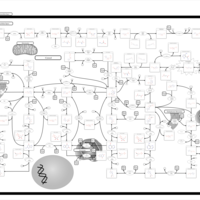 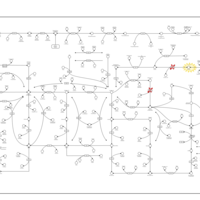 | Not Available |
|
|---|
| Protein Targets |
|---|
| Enzymes | |
| 5'-nucleotidase | NT5E | 6q14-q21 | P21589 | details | | Cytosolic 5'-nucleotidase 1B | NT5C1B | 2p24.2 | Q96P26 | details | | Cytosolic 5'-nucleotidase 1A | NT5C1A | 1p34.3-p33 | Q9BXI3 | details | | 5'(3')-deoxyribonucleotidase, cytosolic type | NT5C | | Q8TCD5 | details | | 5'(3')-deoxyribonucleotidase, mitochondrial | NT5M | | Q9NPB1 | details | | Ectonucleoside triphosphate diphosphohydrolase 1 | ENTPD1 | 10q24 | P49961 | details | | Soluble calcium-activated nucleotidase 1 | CANT1 | 17q25.3 | Q8WVQ1 | details | | Ectonucleoside triphosphate diphosphohydrolase 3 | ENTPD3 | 3p21.3 | O75355 | details | | Inosine triphosphate pyrophosphatase | ITPA | 20p | Q9BY32 | details | | Adenine phosphoribosyltransferase | APRT | 16q24 | P07741 | details | | Hypoxanthine-guanine phosphoribosyltransferase | HPRT1 | Xq26.1 | P00492 | details | | Retinal cone rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit gamma | PDE6H | 12p13 | Q13956 | details | | Calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1A | PDE1A | 2q32.1 | P54750 | details | | High affinity cAMP-specific and IBMX-insensitive 3',5'-cyclic phosphodiesterase 8A | PDE8A | 15q25.3 | O60658 | details | | Calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B | PDE1B | 12q13 | Q01064 | details | | cGMP-inhibited 3',5'-cyclic phosphodiesterase B | PDE3B | 11p15.1 | Q13370 | details | | Cone cGMP-specific 3',5'-cyclic phosphodiesterase subunit alpha' | PDE6C | 10q24 | P51160 | details | | High affinity cAMP-specific 3',5'-cyclic phosphodiesterase 7A | PDE7A | 8q13 | Q13946 | details | | GMP synthase [glutamine-hydrolyzing] | GMPS | 3q24 | P49915 | details | | cGMP-inhibited 3',5'-cyclic phosphodiesterase A | PDE3A | 12p12 | Q14432 | details | | cAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A | PDE10A | 6q26 | Q9Y233 | details | | Calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1C | PDE1C | 7p14.3 | Q14123 | details | | cAMP-specific 3',5'-cyclic phosphodiesterase 4B | PDE4B | 1p31 | Q07343 | details | | cAMP-specific 3',5'-cyclic phosphodiesterase 7B | PDE7B | 6q23-q24 | Q9NP56 | details | | High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A | PDE9A | 21q22.3 | O76083 | details | | cGMP-dependent 3',5'-cyclic phosphodiesterase | PDE2A | 11q13.4 | O00408 | details | | Bis(5'-nucleosyl)-tetraphosphatase [asymmetrical] | NUDT2 | 9p13 | P50583 | details | | Rod cGMP-specific 3',5'-cyclic phosphodiesterase subunit alpha | PDE6A | 5q31.2-q34 | P16499 | details | | High affinity cAMP-specific and IBMX-insensitive 3',5'-cyclic phosphodiesterase 8B | PDE8B | 5q13.3 | O95263 | details | | cAMP-specific 3',5'-cyclic phosphodiesterase 4C | PDE4C | 19p13.11 | Q08493 | details | | Rod cGMP-specific 3',5'-cyclic phosphodiesterase subunit beta | PDE6B | 4p16.3 | P35913 | details | | cAMP-specific 3',5'-cyclic phosphodiesterase 4D | PDE4D | 5q12 | Q08499 | details | | Retinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit gamma | PDE6G | 17q25 | P18545 | details | | Retinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit delta | PDE6D | 2q35-q36 | O43924 | details | | cAMP-specific 3',5'-cyclic phosphodiesterase 4A | PDE4A | 19p13.2 | P27815 | details | | cGMP-specific 3',5'-cyclic phosphodiesterase | PDE5A | 4q27 | O76074 | details | | Protein-glutamine gamma-glutamyltransferase E | TGM3 | 20q11.2 | Q08188 | details | | GMP reductase 2 | GMPR2 | 14q12 | Q9P2T1 | details | | GMP reductase 1 | GMPR | 6p23 | P36959 | details | | Ectonucleoside triphosphate diphosphohydrolase 4 | ENTPD4 | 8p21.3 | Q9Y227 | details | | Ectonucleoside triphosphate diphosphohydrolase 6 | ENTPD6 | 20p11.21 | O75354 | details | | Ectonucleoside triphosphate diphosphohydrolase 5 | ENTPD5 | 14q24 | O75356 | details | | Bifunctional purine biosynthesis protein PURH | ATIC | 2q35 | P31939 | details | | Guanylate kinase | GUK1 | 1q32-q41 | Q16774 | details | | Disks large homolog 4 | DLG4 | 17p13.1 | P78352 | details | | Rho guanine nucleotide exchange factor 1 | ARHGEF1 | 19q13.13 | Q92888 | details | | Guanine nucleotide-binding protein G(s) subunit alpha isoforms short | GNAS | 20q13.3 | P63092 | details | | Vinexin | SORBS3 | 8p21.3 | O60504 | details | | GTPase HRas | HRAS | 11p15.5 | P01112 | details | | Guanine nucleotide-binding protein G(q) subunit alpha | GNAQ | 9q21 | P50148 | details | | Guanine nucleotide-binding protein subunit alpha-11 | GNA11 | 19p13.3 | P29992 | details | | Guanine nucleotide-binding protein G(t) subunit alpha-1 | GNAT1 | 3p21 | P11488 | details | | Protein ALEX | GNAS | 20q13.3 | P84996 | details | | Lysophosphatidic acid receptor 2 | LPAR2 | 19p12 | Q9HBW0 | details | | Cytosolic 5'-nucleotidase 3 | NT5C3 | 7p14.3 | Q9H0P0 | details | | ENTPD4 protein | ENTPD4 | 8p21.3 | Q8NE73 | details | | Cytosolic purine 5'-nucleotidase | NT5C2 | 10q24.32 | P49902 | details | | Dual 3',5'-cyclic-AMP and -GMP phosphodiesterase 11A | PDE11A | 2q31.2 | Q9HCR9 | details | | Ectonucleoside triphosphate diphosphohydrolase 8 | ENTPD8 | 9q34.3 | Q5MY95 | details | | GTPase-activating protein and VPS9 domain-containing protein 1 | GAPVD1 | 9q33.3 | Q14C86 | details | | Probable E3 ubiquitin-protein ligase HERC1 | HERC1 | 15q22 | Q15751 | details | | Rho-related GTP-binding protein RhoU | RHOU | 1q42.11-q42.3 | Q7L0Q8 | details | | Protein RCC2 | RCC2 | 1p36.13 | Q9P258 | details | | Protein very KIND | KNDC1 | 10q26.3 | Q76NI1 | details |
|
|---|
| Transporters | |
| Cyclic nucleotide-gated cation channel alpha-3 | CNGA3 | 2q11.2 | Q16281 | details |
|
|---|
| Metal Bindings | |
| 5'-nucleotidase | NT5E | 6q14-q21 | P21589 | details | | Cytosolic 5'-nucleotidase 1B | NT5C1B | 2p24.2 | Q96P26 | details | | Cytosolic 5'-nucleotidase 1A | NT5C1A | 1p34.3-p33 | Q9BXI3 | details | | 5'(3')-deoxyribonucleotidase, cytosolic type | NT5C | | Q8TCD5 | details | | 5'(3')-deoxyribonucleotidase, mitochondrial | NT5M | | Q9NPB1 | details | | Soluble calcium-activated nucleotidase 1 | CANT1 | 17q25.3 | Q8WVQ1 | details | | Inosine triphosphate pyrophosphatase | ITPA | 20p | Q9BY32 | details | | Calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1A | PDE1A | 2q32.1 | P54750 | details | | High affinity cAMP-specific and IBMX-insensitive 3',5'-cyclic phosphodiesterase 8A | PDE8A | 15q25.3 | O60658 | details | | Calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B | PDE1B | 12q13 | Q01064 | details | | cGMP-inhibited 3',5'-cyclic phosphodiesterase B | PDE3B | 11p15.1 | Q13370 | details | | Cone cGMP-specific 3',5'-cyclic phosphodiesterase subunit alpha' | PDE6C | 10q24 | P51160 | details | | High affinity cAMP-specific 3',5'-cyclic phosphodiesterase 7A | PDE7A | 8q13 | Q13946 | details | | cGMP-inhibited 3',5'-cyclic phosphodiesterase A | PDE3A | 12p12 | Q14432 | details | | cAMP and cAMP-inhibited cGMP 3',5'-cyclic phosphodiesterase 10A | PDE10A | 6q26 | Q9Y233 | details | | Calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1C | PDE1C | 7p14.3 | Q14123 | details | | cAMP-specific 3',5'-cyclic phosphodiesterase 4B | PDE4B | 1p31 | Q07343 | details | | cAMP-specific 3',5'-cyclic phosphodiesterase 7B | PDE7B | 6q23-q24 | Q9NP56 | details | | High affinity cGMP-specific 3',5'-cyclic phosphodiesterase 9A | PDE9A | 21q22.3 | O76083 | details | | cGMP-dependent 3',5'-cyclic phosphodiesterase | PDE2A | 11q13.4 | O00408 | details | | Rod cGMP-specific 3',5'-cyclic phosphodiesterase subunit alpha | PDE6A | 5q31.2-q34 | P16499 | details | | High affinity cAMP-specific and IBMX-insensitive 3',5'-cyclic phosphodiesterase 8B | PDE8B | 5q13.3 | O95263 | details | | cAMP-specific 3',5'-cyclic phosphodiesterase 4C | PDE4C | 19p13.11 | Q08493 | details | | Rod cGMP-specific 3',5'-cyclic phosphodiesterase subunit beta | PDE6B | 4p16.3 | P35913 | details | | cAMP-specific 3',5'-cyclic phosphodiesterase 4D | PDE4D | 5q12 | Q08499 | details | | cAMP-specific 3',5'-cyclic phosphodiesterase 4A | PDE4A | 19p13.2 | P27815 | details | | cGMP-specific 3',5'-cyclic phosphodiesterase | PDE5A | 4q27 | O76074 | details | | GMP reductase 2 | GMPR2 | 14q12 | Q9P2T1 | details | | GMP reductase 1 | GMPR | 6p23 | P36959 | details | | Cytosolic 5'-nucleotidase 3 | NT5C3 | 7p14.3 | Q9H0P0 | details | | Cytosolic purine 5'-nucleotidase | NT5C2 | 10q24.32 | P49902 | details | | Dual 3',5'-cyclic-AMP and -GMP phosphodiesterase 11A | PDE11A | 2q31.2 | Q9HCR9 | details | | Dedicator of cytokinesis protein 2 | DOCK2 | 5q35.1 | Q92608 | details | | Ras guanyl-releasing protein 3 | RASGRP3 | 2p25.1-p24.1 | Q8IV61 | details |
|
|---|
| Receptors | |
| Retinal cone rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit gamma | PDE6H | 12p13 | Q13956 | details | | Calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1A | PDE1A | 2q32.1 | P54750 | details | | Calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1B | PDE1B | 12q13 | Q01064 | details | | cGMP-inhibited 3',5'-cyclic phosphodiesterase B | PDE3B | 11p15.1 | Q13370 | details | | Calcium/calmodulin-dependent 3',5'-cyclic nucleotide phosphodiesterase 1C | PDE1C | 7p14.3 | Q14123 | details | | Retinal rod rhodopsin-sensitive cGMP 3',5'-cyclic phosphodiesterase subunit gamma | PDE6G | 17q25 | P18545 | details | | Atrial natriuretic peptide receptor 2 | NPR2 | 9p21-p12 | P20594 | details | | Atrial natriuretic peptide receptor 1 | NPR1 | 1q21-q22 | P16066 | details | | Heat-stable enterotoxin receptor | GUCY2C | 12p12 | P25092 | details | | Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-2 | GNG2 | 14q21 | P59768 | details | | Guanine nucleotide-binding protein G(s) subunit alpha isoforms short | GNAS | 20q13.3 | P63092 | details | | Guanine nucleotide-binding protein G(q) subunit alpha | GNAQ | 9q21 | P50148 | details | | Guanine nucleotide-binding protein subunit alpha-11 | GNA11 | 19p13.3 | P29992 | details | | Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-11 | GNG11 | 7q21 | P61952 | details | | Guanine nucleotide-binding protein G(t) subunit alpha-1 | GNAT1 | 3p21 | P11488 | details | | Lysophosphatidic acid receptor 2 | LPAR2 | 19p12 | Q9HBW0 | details | | Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-10 | GNG10 | 9q31.3 | P50151 | details | | Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-12 | GNG12 | 1p31.3 | Q9UBI6 | details | | Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-13 | GNG13 | 16p13.3 | Q9P2W3 | details | | Guanine nucleotide-binding protein G(T) subunit gamma-T1 | GNGT1 | 7q21.3 | P63211 | details | | Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-3 | GNG3 | 11p11 | P63215 | details | | Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-4 | GNG4 | 1q42.3 | P50150 | details | | Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-5 | GNG5 | 1p22 | P63218 | details | | Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-7 | GNG7 | 19p13.3 | O60262 | details | | Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-8 | GNG8 | 19q13.32 | Q9UK08 | details | | Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-T2 | GNGT2 | 17q21 | O14610 | details | | T-lymphoma invasion and metastasis-inducing protein 2 | TIAM2 | 6q25.2 | Q8IVF5 | details | | Guanine nucleotide-binding protein G(i) subunit alpha-1 | GNAI1 | 7q21 | P63096 | details | | Guanine nucleotide-binding protein G(s) subunit alpha isoforms XLas | GNAS | 20q13.3 | Q5JWF2 | details | | Guanine nucleotide-binding protein subunit alpha-14 | GNA14 | 9q21 | O95837 | details | | Guanine nucleotide-binding protein G(k) subunit alpha | GNAI3 | 1p13 | P08754 | details | | Guanine nucleotide-binding protein G(i) subunit alpha-2 | GNAI2 | 3p21 | P04899 | details | | Guanine nucleotide-binding protein G(t) subunit alpha-2 | GNAT2 | 1p13.1 | P19087 | details | | Guanine nucleotide-binding protein G(o) subunit alpha | GNAO1 | 16q13 | P09471 | details | | Guanine nucleotide-binding protein subunit alpha-13 | GNA13 | 17q24.3 | Q14344 | details | | Guanine nucleotide binding protein (G protein), alpha 14 | GNA14 | 9q21 | B1ALW3 | details | | Guanine nucleotide-binding protein subunit alpha-12 | GNA12 | 7p22.2 | Q03113 | details | | Growth factor receptor-bound protein 2 | GRB2 | 17q24-q25 | P62993 | details | | Guanine nucleotide binding protein (G protein), q polypeptide | GNAQ | 9q21 | B1AM21 | details | | Guanine nucleotide-binding protein G(z) subunit alpha | GNAZ | 22q11.22 | P19086 | details | | Guanine nucleotide-binding protein G(olf) subunit alpha | GNAL | 18p11.22-p11.21 | P38405 | details | | Guanine nucleotide-binding protein subunit alpha-15 | GNA15 | 19p13.3 | P30679 | details |
|
|---|
| Transcriptional Factors | |
| High affinity cAMP-specific and IBMX-insensitive 3',5'-cyclic phosphodiesterase 8A | PDE8A | 15q25.3 | O60658 | details | | cGMP-dependent 3',5'-cyclic phosphodiesterase | PDE2A | 11q13.4 | O00408 | details | | High affinity cAMP-specific and IBMX-insensitive 3',5'-cyclic phosphodiesterase 8B | PDE8B | 5q13.3 | O95263 | details |
|
|---|
| Concentrations Data |
|---|
| Not Available |
|---|
| External Links |
|---|
| HMDB ID | HMDB0001397 |
|---|
| DrugBank ID | DB01972 |
|---|
| Phenol Explorer Compound ID | Not Available |
|---|
| FoodDB ID | FDB030896 |
|---|
| KNApSAcK ID | C00019635 |
|---|
| Chemspider ID | 6545 |
|---|
| KEGG Compound ID | C00144 |
|---|
| BioCyc ID | GMP |
|---|
| BiGG ID | 34024 |
|---|
| Wikipedia Link | Guanosine_monophosphate |
|---|
| METLIN ID | 6216 |
|---|
| PubChem Compound | 6804 |
|---|
| PDB ID | Not Available |
|---|
| ChEBI ID | 17345 |
|---|
| References |
|---|
| General References | Not Available |
|---|

