| Record Information |
|---|
| Version | 1.0 |
|---|
| Created at | 2020-04-17 18:44:58 UTC |
|---|
| Updated at | 2020-11-18 16:38:53 UTC |
|---|
| CannabisDB ID | CDB004867 |
|---|
| Secondary Accession Numbers | Not Available |
|---|
| Cannabis Compound Identification |
|---|
| Common Name | Nicotinamide riboside |
|---|
| Description | Nicotinamide riboside, also known as N-ribosylnicotinamide or SRT-647, belongs to the class of organic compounds known as glycosylamines. Glycosylamines are compounds consisting of an amine with a beta-N-glycosidic bond to a carbohydrate, thus forming a cyclic hemiaminal ether bond (alpha-amino ether). A pyridine nucleoside consisting of nicotinamide with a beta-D-ribofuranosyl moiety at the 1-position. Nicotinamide riboside is an extremely weak basic (essentially neutral) compound (based on its pKa). Nicotinamide riboside exists in all living species, ranging from bacteria to humans. Within humans, nicotinamide riboside participates in a number of enzymatic reactions. In particular, nicotinamide riboside and phosphoric acid can be biosynthesized from niacinamide and ribose-1-arsenate through its interaction with the enzyme purine nucleoside phosphorylase. In addition, nicotinamide riboside can be converted into nicotinamide ribotide through its interaction with the enzyme cytosolic purine 5'-nucleotidase. In humans, nicotinamide riboside is involved in nicotinate and nicotinamide metabolism. Outside of the human body, Nicotinamide riboside has been detected, but not quantified in, milk (cow). This could make nicotinamide riboside a potential biomarker for the consumption of these foods. Nicotinamide riboside is expected to be in Cannabis as all living plants are known to produce and metabolize it. |
|---|
| Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| 1-(beta-D-Ribofuranosyl)nicotinamide | ChEBI | | beta-Nicotinamide D-riboside | ChEBI | | Nicotinamide ribonucleoside | ChEBI | | Nicotinamide ribose | ChEBI | | Nicotinamide-beta-riboside | ChEBI | | N-Ribosylnicotinamide | Kegg | | 1-(b-D-Ribofuranosyl)nicotinamide | Generator | | 1-(Β-D-ribofuranosyl)nicotinamide | Generator | | b-Nicotinamide D-riboside | Generator | | Β-nicotinamide D-riboside | Generator | | Nicotinamide-b-riboside | Generator | | Nicotinamide-β-riboside | Generator | | 1-b-D-Ribosyl-3-pyridinecarboxamide | HMDB | | 1-beta-D-Ribosyl-3-pyridinecarboxamide | HMDB | | 1-beta-delta-Ribosyl-3-pyridinecarboxamide | HMDB | | 3-(Aminocarbonyl)-1-beta-D-ribofuranosyl-pyridinium | HMDB | | 3-(Aminocarbonyl)-1-beta-delta-ribofuranosyl-pyridinium | HMDB | | Ribosylnicotinamide | HMDB | | SRT-647 | HMDB | | SRT 647 | HMDB |
|
|---|
| Chemical Formula | C11H15N2O5 |
|---|
| Average Molecular Weight | 255.25 |
|---|
| Monoisotopic Molecular Weight | 255.0981 |
|---|
| IUPAC Name | 3-carbamoyl-1-[(2R,3R,4S,5R)-3,4-dihydroxy-5-(hydroxymethyl)oxolan-2-yl]-1lambda5-pyridin-1-ylium |
|---|
| Traditional Name | 3-carbamoyl-1-[(2R,3R,4S,5R)-3,4-dihydroxy-5-(hydroxymethyl)oxolan-2-yl]-1lambda5-pyridin-1-ylium |
|---|
| CAS Registry Number | 1341-23-7 |
|---|
| SMILES | NC(=O)C1=C[N+](=CC=C1)[C@@H]1O[C@H](CO)[C@@H](O)[C@H]1O |
|---|
| InChI Identifier | InChI=1S/C11H14N2O5/c12-10(17)6-2-1-3-13(4-6)11-9(16)8(15)7(5-14)18-11/h1-4,7-9,11,14-16H,5H2,(H-,12,17)/p+1/t7-,8-,9-,11-/m1/s1 |
|---|
| InChI Key | JLEBZPBDRKPWTD-TURQNECASA-O |
|---|
| Chemical Taxonomy |
|---|
| Description | Belongs to the class of organic compounds known as glycosylamines. Glycosylamines are compounds consisting of an amine with a beta-N-glycosidic bond to a carbohydrate, thus forming a cyclic hemiaminal ether bond (alpha-amino ether). |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic oxygen compounds |
|---|
| Class | Organooxygen compounds |
|---|
| Sub Class | Carbohydrates and carbohydrate conjugates |
|---|
| Direct Parent | Glycosylamines |
|---|
| Alternative Parents | |
|---|
| Substituents | - N-glycosyl compound
- Pentose monosaccharide
- Nicotinamide
- Pyridine carboxylic acid or derivatives
- Monosaccharide
- Pyridine
- Pyridinium
- Vinylogous amide
- Heteroaromatic compound
- Tetrahydrofuran
- Secondary alcohol
- Carboxamide group
- Primary carboxylic acid amide
- Organoheterocyclic compound
- Oxacycle
- Azacycle
- Carboxylic acid derivative
- Hydrocarbon derivative
- Primary alcohol
- Organic oxide
- Organonitrogen compound
- Organopnictogen compound
- Alcohol
- Organic nitrogen compound
- Organic cation
- Aromatic heteromonocyclic compound
|
|---|
| Molecular Framework | Aromatic heteromonocyclic compounds |
|---|
| External Descriptors | |
|---|
| Ontology |
|---|
|
| Disposition | Source: Biological location: |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Experimental Properties | | Property | Value | Reference |
|---|
| Melting Point | Not Available | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | Not Available | Not Available | | logP | Not Available | Not Available |
|
|---|
| Predicted Properties | [] |
|---|
| Spectra |
|---|
| EI-MS/GC-MS | | Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | Nicotinamide riboside, non-derivatized, Predicted GC-MS Spectrum - 70eV, Positive | splash10-070f-9530000000-71d9e02ee640bb800185 | Spectrum | | Predicted GC-MS | Nicotinamide riboside, 3 TMS, Predicted GC-MS Spectrum - 70eV, Positive | splash10-0a6r-9825400000-99de0fc632f408e32a12 | Spectrum | | Predicted GC-MS | Nicotinamide riboside, non-derivatized, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Nicotinamide riboside, TMS_1_1, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Nicotinamide riboside, TMS_1_2, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Nicotinamide riboside, TMS_1_3, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Nicotinamide riboside, TMS_1_4, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Nicotinamide riboside, TMS_2_1, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Nicotinamide riboside, TMS_2_2, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Nicotinamide riboside, TMS_2_3, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Nicotinamide riboside, TMS_2_4, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Nicotinamide riboside, TMS_2_5, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Nicotinamide riboside, TMS_2_6, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Nicotinamide riboside, TMS_2_7, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Nicotinamide riboside, TMS_3_2, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Nicotinamide riboside, TMS_3_3, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Nicotinamide riboside, TMS_3_4, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Nicotinamide riboside, TMS_3_5, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Nicotinamide riboside, TMS_3_6, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Nicotinamide riboside, TMS_3_7, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Nicotinamide riboside, TBDMS_1_1, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Nicotinamide riboside, TBDMS_1_2, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Nicotinamide riboside, TBDMS_1_3, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Nicotinamide riboside, TBDMS_1_4, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | Nicotinamide riboside, TBDMS_2_1, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum |
|
|---|
| MS/MS | | Type | Description | Splash Key | View |
|---|
| MS/MS | LC-MS/MS Spectrum - Orbitrap 0V, positive | splash10-0a4i-0190000000-51bf764225f08b566d36 | 2020-07-22 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - Orbitrap 1V, positive | splash10-0a4i-0290000000-ffbe280293f874204044 | 2020-07-22 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - Orbitrap 2V, positive | splash10-05fr-0970000000-1714e669425e34d3754a | 2020-07-22 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - Orbitrap 3V, positive | splash10-00di-0920000000-77ef12950b3152d752c8 | 2020-07-22 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - Orbitrap 5V, positive | splash10-00di-0900000000-b14d15851532222ba74a | 2020-07-22 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - Orbitrap 6V, positive | splash10-00di-0900000000-277d95be8f0485eb3ce5 | 2020-07-22 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - Orbitrap 7V, positive | splash10-00di-0900000000-354c1f3978635e5109bb | 2020-07-22 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - Orbitrap 8V, positive | splash10-00di-0900000000-ea5e376f71c6414964bc | 2020-07-22 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - Orbitrap 10V, positive | splash10-00di-0900000000-6b289ae1c5f53e92f774 | 2020-07-22 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - Orbitrap 11V, positive | splash10-00di-1900000000-6ad92fa16e2cca17f959 | 2020-07-22 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - Orbitrap 12V, positive | splash10-00di-1900000000-cee8757ed6b406ccf044 | 2020-07-22 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - Orbitrap 15V, positive | splash10-00di-2900000000-78f21d967892a0d9ab85 | 2020-07-22 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - Orbitrap 19V, positive | splash10-00di-3900000000-cd1f66bfc8ded7bd098e | 2020-07-22 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - Orbitrap 22V, positive | splash10-00e9-8900000000-5a6bc8100043575c6a58 | 2020-07-22 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - Orbitrap 28V, positive | splash10-00ai-9200000000-aedd3c5bf179b666a6ae | 2020-07-22 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - Orbitrap 33V, positive | splash10-003r-9000000000-22053c801abdfaf5f562 | 2020-07-22 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - Orbitrap 41V, positive | splash10-0fai-9000000000-b6b4b7e819e08c863a9a | 2020-07-22 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - n/a 17V, positive | splash10-00di-0900000000-3364e345334aa7f5a742 | 2020-07-22 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - n/a 17V, positive | splash10-004i-9000000000-3bf262f96423e85a1756 | 2020-07-22 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0a4i-0090000000-e25d1beb47cb1fe9e4cb | 2015-09-15 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-002b-3090000000-03687e9ef697eda1d6c8 | 2015-09-15 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-052g-9300000000-2755bde4df843d2227f2 | 2015-09-15 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0udi-0090000000-809be1146b2a2c8d96a7 | 2015-09-15 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0udi-1390000000-2fc72cf0619f182e044f | 2015-09-15 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0k96-9300000000-4d5b96b192f3484b1b8a | 2015-09-15 | View Spectrum |
|
|---|
| NMR | Not Available |
|---|
| Pathways |
|---|
| Pathways | | Name | SMPDB/Pathwhiz | KEGG | | Nicotinate and Nicotinamide Metabolism | 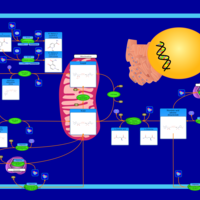 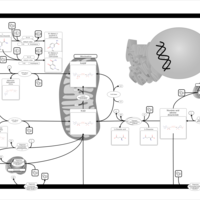  | 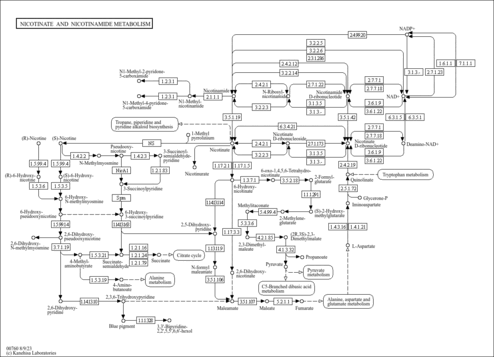 |
|
|---|
| Protein Targets |
|---|
| Enzymes | |
|---|
| Transporters | Not Available |
|---|
| Metal Bindings | |
|---|
| Receptors | Not Available |
|---|
| Transcriptional Factors | Not Available |
|---|
| Concentrations Data |
|---|
| Not Available |
|---|
| External Links |
|---|
| HMDB ID | HMDB0000855 |
|---|
| DrugBank ID | Not Available |
|---|
| Phenol Explorer Compound ID | Not Available |
|---|
| FoodDB ID | FDB022281 |
|---|
| KNApSAcK ID | Not Available |
|---|
| Chemspider ID | 388956 |
|---|
| KEGG Compound ID | C03150 |
|---|
| BioCyc ID | NICOTINAMIDE_RIBOSE |
|---|
| BiGG ID | 41300 |
|---|
| Wikipedia Link | Nicotinamide riboside |
|---|
| METLIN ID | 5818 |
|---|
| PubChem Compound | 439924 |
|---|
| PDB ID | Not Available |
|---|
| ChEBI ID | 15927 |
|---|
| References |
|---|
| General References | Not Available |
|---|
