| Record Information |
|---|
| Version | 1.0 |
|---|
| Created at | 2020-04-17 18:46:02 UTC |
|---|
| Updated at | 2020-11-18 16:38:55 UTC |
|---|
| CannabisDB ID | CDB004877 |
|---|
| Secondary Accession Numbers | Not Available |
|---|
| Cannabis Compound Identification |
|---|
| Common Name | 2'-Deoxyguanosine 5'-monophosphate |
|---|
| Description | 2'-Deoxyguanosine 5'-monophosphate, also known as deoxyguanylic acid or 2'-deoxy-GMP, belongs to the class of organic compounds known as purine 2'-deoxyribonucleoside monophosphates. These are purine nucleotides with monophosphate group linked to the ribose moiety lacking a hydroxyl group at position 2. 2'-Deoxyguanosine 5'-monophosphate is a moderately basic compound (based on its pKa). 2'-Deoxyguanosine 5'-monophosphate exists in all living species, ranging from bacteria to humans. Within humans, 2'-deoxyguanosine 5'-monophosphate participates in a number of enzymatic reactions. In particular, 2'-deoxyguanosine 5'-monophosphate can be converted into dGDP; which is mediated by the enzyme guanylate kinase. In addition, 2'-deoxyguanosine 5'-monophosphate can be converted into deoxyguanosine through its interaction with the enzyme cytosolic purine 5'-nucleotidase. In humans, 2'-deoxyguanosine 5'-monophosphate is involved in the metabolic disorder called the gout or kelley-seegmiller syndrome pathway. A purine 2'-deoxyribonucleoside 5'-monophosphate having guanine as the nucleobase. 2'-Deoxyguanosine 5'-monophosphate is expected to be in Cannabis as all living plants are known to produce and metabolize it. |
|---|
| Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| 2'-Deoxy-GMP | ChEBI | | 2'-Deoxyguanosine 5'-(dihydrogen phosphate) | ChEBI | | 2'-Deoxyguanosine 5'-phosphate | ChEBI | | 2'-Deoxyguanylic acid | ChEBI | | 2'-dGMP | ChEBI | | Deoxy-GMP | ChEBI | | Deoxyguanosine 5'-monophosphate | ChEBI | | Deoxyguanosine 5'-phosphate | ChEBI | | Deoxyguanosine monophosphate | ChEBI | | Deoxyguanylic acid | ChEBI | | dGMP | ChEBI | | 2'-Deoxyguanosine 5'-(dihydrogen phosphoric acid) | Generator | | 2'-Deoxyguanosine 5'-phosphoric acid | Generator | | 2'-Deoxyguanylate | Generator | | Deoxyguanosine 5'-monophosphoric acid | Generator | | Deoxyguanosine 5'-phosphoric acid | Generator | | Deoxyguanosine monophosphoric acid | Generator | | Deoxyguanylate | Generator | | 2'-Deoxyguanosine 5'-monophosphoric acid | Generator | | 2'-Deoxy-5'-GMP | HMDB | | 2'-Deoxy-5'-guanylate | HMDB | | 2'-Deoxy-5'-guanylic acid | HMDB | | 2'-Deoxy-guanosine 5'-(dihydrogen phosphate) | HMDB | | 2'-Deoxy-guanosine 5'-phosphate | HMDB | | 2'-Deoxy-guanosine phosphate | HMDB | | 2'-Deoxyguanosine-5'-phosphate | HMDB | | 2'-DG-5'-MP | HMDB | | Deoxyguanosine-phosphate | HMDB | | Guanine riboside | HMDB | | 2'-Deoxyguanosine 5'-phosphate, ion (1+) | HMDB | | 2'-Deoxyguanosine 5'-phosphate, disodium salt | HMDB |
|
|---|
| Chemical Formula | C10H14N5O7P |
|---|
| Average Molecular Weight | 347.22 |
|---|
| Monoisotopic Molecular Weight | 347.0631 |
|---|
| IUPAC Name | {[(2R,3S,5R)-5-(2-amino-6-oxo-6,9-dihydro-3H-purin-9-yl)-3-hydroxyoxolan-2-yl]methoxy}phosphonic acid |
|---|
| Traditional Name | deoxyguanylate |
|---|
| CAS Registry Number | 902-04-5 |
|---|
| SMILES | NC1=NC2=C(N=CN2[C@H]2C[C@H](O)[C@@H](COP(O)(O)=O)O2)C(=O)N1 |
|---|
| InChI Identifier | InChI=1S/C10H14N5O7P/c11-10-13-8-7(9(17)14-10)12-3-15(8)6-1-4(16)5(22-6)2-21-23(18,19)20/h3-6,16H,1-2H2,(H2,18,19,20)(H3,11,13,14,17)/t4-,5+,6+/m0/s1 |
|---|
| InChI Key | LTFMZDNNPPEQNG-KVQBGUIXSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | Belongs to the class of organic compounds known as purine 2'-deoxyribonucleoside monophosphates. These are purine nucleotides with monophosphate group linked to the ribose moiety lacking a hydroxyl group at position 2. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Nucleosides, nucleotides, and analogues |
|---|
| Class | Purine nucleotides |
|---|
| Sub Class | Purine deoxyribonucleotides |
|---|
| Direct Parent | Purine 2'-deoxyribonucleoside monophosphates |
|---|
| Alternative Parents | |
|---|
| Substituents | - Purine 2'-deoxyribonucleoside monophosphate
- Imidazopyrimidine
- Purine
- Hydroxypyrimidine
- Monoalkyl phosphate
- N-substituted imidazole
- Organic phosphoric acid derivative
- Phosphoric acid ester
- Alkyl phosphate
- Pyrimidine
- Azole
- Imidazole
- Heteroaromatic compound
- Tetrahydrofuran
- Secondary alcohol
- Organoheterocyclic compound
- Azacycle
- Oxacycle
- Organooxygen compound
- Organonitrogen compound
- Hydrocarbon derivative
- Organic oxide
- Organopnictogen compound
- Organic oxygen compound
- Organic nitrogen compound
- Alcohol
- Aromatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aromatic heteropolycyclic compounds |
|---|
| External Descriptors | |
|---|
| Ontology |
|---|
|
| Disposition | Source: Biological location: |
|---|
| Role | Industrial application: |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Experimental Properties | | Property | Value | Reference |
|---|
| Melting Point | Not Available | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | Not Available | Not Available | | logP | Not Available | Not Available |
|
|---|
| Predicted Properties | [] |
|---|
| Spectra |
|---|
| EI-MS/GC-MS | | Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | 2'-Deoxyguanosine 5'-monophosphate, non-derivatized, Predicted GC-MS Spectrum - 70eV, Positive | splash10-0002-9412000000-b0aa92c3ffa6b5b0890c | Spectrum | | Predicted GC-MS | 2'-Deoxyguanosine 5'-monophosphate, 1 TMS, Predicted GC-MS Spectrum - 70eV, Positive | splash10-03dm-9422100000-bc551a9549f9ae975b71 | Spectrum | | Predicted GC-MS | 2'-Deoxyguanosine 5'-monophosphate, non-derivatized, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum |
|
|---|
| MS/MS | | Type | Description | Splash Key | View |
|---|
| MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 10V, Positive (Annotated) | splash10-0fr2-0598000000-21b063e2aff7146d8568 | 2012-07-24 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF (UPLC Q-Tof Premier, Waters) , Positive | splash10-0udi-0900000000-d2858e957ac8a4c91e79 | 2012-08-31 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF (UPLC Q-Tof Premier, Waters) , Negative | splash10-004j-9203000000-40bcf8d5e1ae6813c4f7 | 2012-08-31 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , negative | splash10-004j-9203000000-40bcf8d5e1ae6813c4f7 | 2017-09-14 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - LC-ESI-QTOF , positive | splash10-0udi-0900000000-dab30b9a6de3477fa70e | 2017-09-14 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - 10V, Negative | splash10-002b-6109000000-6e5f13e1030d907bf450 | 2021-09-20 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - 20V, Negative | splash10-004i-9200000000-c3273057674f4f74157b | 2021-09-20 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - 10V, Negative | splash10-002b-6009000000-303a1b69aa5c35e59b53 | 2021-09-20 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - 40V, Negative | splash10-004i-9200000000-5a760c04146b7ec031d7 | 2021-09-20 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - 40V, Negative | splash10-004i-9300000000-d435ed7c5cdb98529682 | 2021-09-20 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - 20V, Negative | splash10-004i-9100000000-41c0a9342f54a46fa4e4 | 2021-09-20 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - 40V, Positive | splash10-000i-2900000000-68f72700dfdbb0374a85 | 2021-09-20 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - 10V, Positive | splash10-000j-0905000000-26458e3261a5441d9304 | 2021-09-20 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - 40V, Positive | splash10-0udr-1900000000-7a7e7e8e0b0145b7230d | 2021-09-20 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - 20V, Positive | splash10-0udi-1900000000-a0ce6815866f122bcce0 | 2021-09-20 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - 10V, Positive | splash10-000i-0901000000-415ae3ba37bc56d4a402 | 2021-09-20 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - 20V, Positive | splash10-000i-1900000000-bd375d50d1fb3f236468 | 2021-09-20 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - 10V, Positive | splash10-0udi-0900000000-6585941fd6af257d9a96 | 2021-09-20 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - 40V, Positive | splash10-000i-2900000000-303e1f2bb06730d8520d | 2021-09-20 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0udi-0901000000-e4234086a26601ce5af3 | 2016-09-12 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0udi-0900000000-566d720734de1bad5cd6 | 2016-09-12 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0udi-0900000000-5d4689698c352d2a2f45 | 2016-09-12 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-002b-6309000000-10f0b25fe6368edd1660 | 2016-09-12 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-004i-9500000000-d531006102bbcb9090d1 | 2016-09-12 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-004i-9100000000-a08be881b0ccce87df8a | 2016-09-12 | View Spectrum |
|
|---|
| NMR | | Type | Description | | View |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 600 MHz, H2O, experimental) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, D2O, experimental) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 100 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 100 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 1000 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 200 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 200 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 300 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 300 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 400 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 500 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 500 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 600 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 600 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 700 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 700 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 800 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 800 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 900 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 900 MHz, D2O, predicted) | | Spectrum | | 2D NMR | [1H, 13C]-HSQC NMR Spectrum (2D, 600 MHz, H2O, experimental) | | Spectrum |
|
|---|
| Pathways |
|---|
| Pathways | | Name | SMPDB/Pathwhiz | KEGG | | Purine Metabolism |  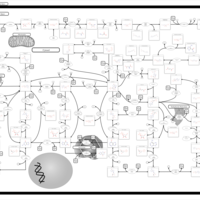 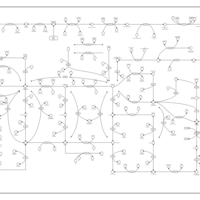 |  | | Adenosine Deaminase Deficiency |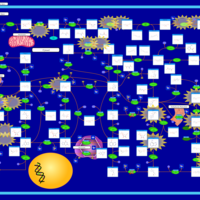  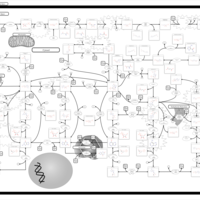 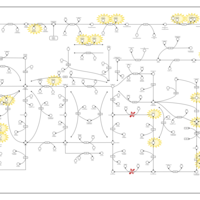 | Not Available | | Adenylosuccinate Lyase Deficiency | 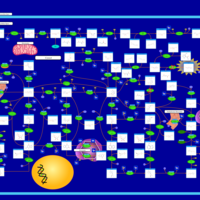 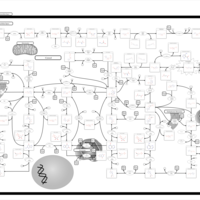 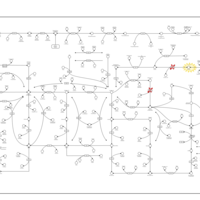 | Not Available | | Gout or Kelley-Seegmiller Syndrome | 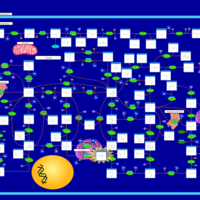 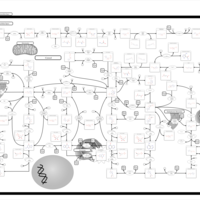 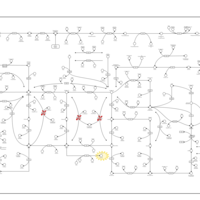 | Not Available | | Lesch-Nyhan Syndrome (LNS) | 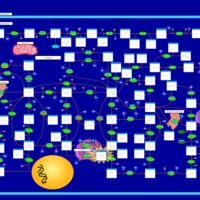 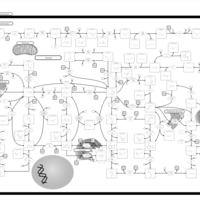 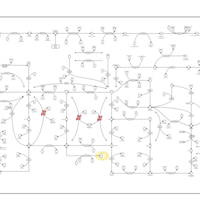 | Not Available |
|
|---|
| Protein Targets |
|---|
| Enzymes | |
|---|
| Transporters | Not Available |
|---|
| Metal Bindings | |
|---|
| Receptors | Not Available |
|---|
| Transcriptional Factors | Not Available |
|---|
| Concentrations Data |
|---|
| Not Available |
|---|
| External Links |
|---|
| HMDB ID | HMDB0001044 |
|---|
| DrugBank ID | DB04457 |
|---|
| Phenol Explorer Compound ID | Not Available |
|---|
| FoodDB ID | FDB022388 |
|---|
| KNApSAcK ID | C00019356 |
|---|
| Chemspider ID | 58570 |
|---|
| KEGG Compound ID | C00362 |
|---|
| BioCyc ID | DGMP |
|---|
| BiGG ID | 34744 |
|---|
| Wikipedia Link | Deoxyguanosine_monophosphate |
|---|
| METLIN ID | Not Available |
|---|
| PubChem Compound | 65059 |
|---|
| PDB ID | Not Available |
|---|
| ChEBI ID | 16192 |
|---|
| References |
|---|
| General References | Not Available |
|---|
