| Record Information |
|---|
| Version | 1.0 |
|---|
| Created at | 2020-03-30 18:00:52 UTC |
|---|
| Updated at | 2020-12-07 19:07:55 UTC |
|---|
| CannabisDB ID | CDB001059 |
|---|
| Secondary Accession Numbers | Not Available |
|---|
| Cannabis Compound Identification |
|---|
| Common Name | PE(16:0/20:0) |
|---|
| Description | PE(16:0/20:0) is a phosphatidylethanolamine. It is a glycerophospholipid in which a phosphorylethanolamine moiety occupies a glycerol substitution site. As is the case with diacylglycerols, glycerophosphoethanolamines can have many different combinations of fatty acids of varying lengths and saturation attached to the C-1 and C-2 atoms. PE(16:0/20:0), in particular, consists of one hexadecanoyl chain to the C-1 atom, and one eicosanoyl to the C-2 atom. While most phospholipids have a saturated fatty acid on C-1 and an unsaturated fatty acid on C-2 of the glycerol backbone, the fatty acid distribution at the C-1 and C-2 positions of glycerol within phospholipids is continually in flux, owing to phospholipid degradation and the continuous phospholipid remodeling that occurs while these molecules are in membranes. PEs are neutral zwitterions at physiological pH. They mostly have palmitic or stearic acid on carbon 1 and a long chain unsaturated fatty acid (e.g. 18:2, 20:4 and 22:6) on carbon 2. PE synthesis can occur via two pathways. The first requires that ethanolamine be activated by phosphorylation and then coupled to CDP. The ethanolamine is then transferred from CDP-ethanolamine to phosphatidic acid to yield PE. The second involves the decarboxylation of PS. This compound is expected to be in Cannabis as all living plants are known to produce and metabolize it. |
|---|
| Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| (2-Aminoethoxy)[(2R)-3-(hexadecanoyloxy)-2-(icosanoyloxy)propoxy]phosphinic acid | ChEBI | | 1-Hexadecanoyl-2-eicosanoyl-sn-glycero-3-phosphoethanolamine | ChEBI | | 1-Palmitoyl-2-arachidoyl-sn-glycero-3-phosphoethanolamine | ChEBI | | PE(36:0) | ChEBI | | Phosphatidylethanolamine(16:0/20:0) | ChEBI | | Phosphatidylethanolamine(36:0) | ChEBI | | (2-Aminoethoxy)[(2R)-3-(hexadecanoyloxy)-2-(icosanoyloxy)propoxy]phosphinate | Generator | | Phophatidylethanolamine(36:0) | HMDB | | 1-Palmitoyl-2-arachidonyl-sn-glycero-3-phosphoethanolamine | HMDB | | Phophatidylethanolamine(16:0/20:0) | HMDB | | GPEtn(36:0) | HMDB | | GPEtn(16:0/20:0) | HMDB | | PE(16:0/20:0) | Lipid Annotator |
|
|---|
| Chemical Formula | C41H82NO8P |
|---|
| Average Molecular Weight | 748.07 |
|---|
| Monoisotopic Molecular Weight | 747.5778 |
|---|
| IUPAC Name | (2-aminoethoxy)[(2R)-3-(hexadecanoyloxy)-2-(icosanoyloxy)propoxy]phosphinic acid |
|---|
| Traditional Name | 2-aminoethoxy((2R)-3-(hexadecanoyloxy)-2-(icosanoyloxy)propoxy)phosphinic acid |
|---|
| CAS Registry Number | Not Available |
|---|
| SMILES | [H][C@@](COC(=O)CCCCCCCCCCCCCCC)(COP(O)(=O)OCCN)OC(=O)CCCCCCCCCCCCCCCCCCC |
|---|
| InChI Identifier | InChI=1S/C41H82NO8P/c1-3-5-7-9-11-13-15-17-18-19-20-22-24-26-28-30-32-34-41(44)50-39(38-49-51(45,46)48-36-35-42)37-47-40(43)33-31-29-27-25-23-21-16-14-12-10-8-6-4-2/h39H,3-38,42H2,1-2H3,(H,45,46)/t39-/m1/s1 |
|---|
| InChI Key | ZFCOGJGXPSVFIM-LDLOPFEMSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | Belongs to the class of organic compounds known as phosphatidylethanolamines. These are glycerophosphoetahnolamines in which two fatty acids are bonded to the glycerol moiety through ester linkages. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Glycerophospholipids |
|---|
| Sub Class | Glycerophosphoethanolamines |
|---|
| Direct Parent | Phosphatidylethanolamines |
|---|
| Alternative Parents | |
|---|
| Substituents | - Diacylglycero-3-phosphoethanolamine
- Phosphoethanolamine
- Fatty acid ester
- Dialkyl phosphate
- Dicarboxylic acid or derivatives
- Organic phosphoric acid derivative
- Phosphoric acid ester
- Alkyl phosphate
- Fatty acyl
- Amino acid or derivatives
- Carboxylic acid ester
- Carboxylic acid derivative
- Organopnictogen compound
- Organic oxygen compound
- Organooxygen compound
- Organonitrogen compound
- Amine
- Primary aliphatic amine
- Organic nitrogen compound
- Primary amine
- Carbonyl group
- Organic oxide
- Hydrocarbon derivative
- Aliphatic acyclic compound
|
|---|
| Molecular Framework | Aliphatic acyclic compounds |
|---|
| External Descriptors | |
|---|
| Ontology |
|---|
|
| Physiological effect | Organoleptic effect: |
|---|
| Disposition | Route of exposure: Source: Biological location: |
|---|
| Role | Industrial application: Biological role: |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Experimental Properties | | Property | Value | Reference |
|---|
| Melting Point | Not Available | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | Not Available | Not Available | | logP | Not Available | Not Available |
|
|---|
| Predicted Properties | [] |
|---|
| Spectra |
|---|
| EI-MS/GC-MS | | Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | PE(16:0/20:0), non-derivatized, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum |
|
|---|
| MS/MS | | Type | Description | Splash Key | View |
|---|
| Predicted MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0002-0011000900-9c2dfc58e07043346b91 | 2017-10-04 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0002-0011000900-9c2dfc58e07043346b91 | 2017-10-04 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0bta-0399410600-3e20efb1a79fd27e7ecb | 2017-10-04 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0002-0000001900-4a7b6d9013d36f909f90 | 2017-10-04 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0a4j-0003419700-cea891efa4464a3273fb | 2017-10-04 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0a4i-0003419300-7540ae55bc0161c16f68 | 2017-10-04 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-00di-0000001900-50a418c4f44c815f6289 | 2021-09-22 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-00fr-0000001900-34619ad7f9118f8f3038 | 2021-09-22 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-004i-0100201900-e301490f80913b96997a | 2021-09-22 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-0002-0000001900-bbbbfd5d071fd445575f | 2021-09-23 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-0a4j-0003419700-15c31fc9a5d375ff82d1 | 2021-09-23 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0a4i-0003419300-e201d537cb760e8f8142 | 2021-09-23 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0002-0011000900-bdc842ad36789130e638 | 2021-09-24 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0002-0011000900-bdc842ad36789130e638 | 2021-09-24 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0bta-0399410600-08bc4f59acac111161dc | 2021-09-24 | View Spectrum |
|
|---|
| NMR | Not Available |
|---|
| Pathways |
|---|
| Pathways | | Name | SMPDB/Pathwhiz | KEGG | | Phosphatidylcholine Biosynthesis PC(16:0/20:0) | 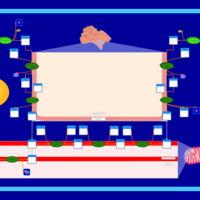 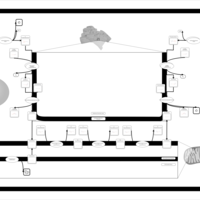 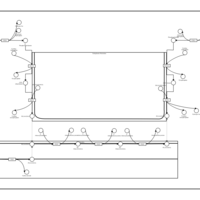 | Not Available | | Phosphatidylethanolamine Biosynthesis PE(16:0/20:0) | 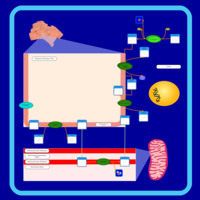 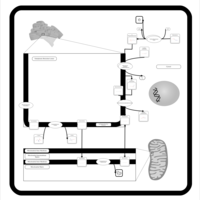 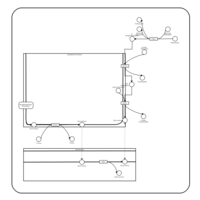 | Not Available |
|
|---|
| Protein Targets |
|---|
| Enzymes | |
|---|
| Transporters | |
| Probable phospholipid-transporting ATPase IG | ATP11C | | Q8NB49 | details | | Probable phospholipid-transporting ATPase IH | ATP11A | 13q34 | P98196 | details | | Probable phospholipid-transporting ATPase VA | ATP10A | 15q11.2 | O60312 | details | | Probable phospholipid-transporting ATPase IC | ATP8B1 | 18q21-q22|18q21.31 | O43520 | details | | Probable phospholipid-transporting ATPase IIA | ATP9A | 20q13.2 | O75110 | details | | Probable phospholipid-transporting ATPase VD | ATP10D | 4p12 | Q9P241 | details | | Probable phospholipid-transporting ATPase IB | ATP8A2 | 13q12 | Q9NTI2 | details | | Probable phospholipid-transporting ATPase IA | ATP8A1 | 4p13 | Q9Y2Q0 | details | | Probable phospholipid-transporting ATPase IM | ATP8B4 | 15q21.2 | Q8TF62 | details | | Probable phospholipid-transporting ATPase IF | ATP11B | 3q27 | Q9Y2G3 | details | | Probable phospholipid-transporting ATPase IK | ATP8B3 | 19p13.3 | O60423 | details | | Probable phospholipid-transporting ATPase VB | ATP10B | 5q34 | O94823 | details | | Probable phospholipid-transporting ATPase ID | ATP8B2 | 1q21.3 | P98198 | details |
|
|---|
| Metal Bindings | |
| Calcium-dependent phospholipase A2 | PLA2G5 | 1p36-p34 | P39877 | details | | Group IIF secretory phospholipase A2 | PLA2G2F | 1p35 | Q9BZM2 | details | | Phospholipase A2 | PLA2G1B | 12q23-q24.1 | P04054 | details | | Group XIIB secretory phospholipase A2-like protein | PLA2G12B | | Q9BX93 | details | | Group 10 secretory phospholipase A2 | PLA2G10 | 16p13.1-p12 | O15496 | details | | Group IIE secretory phospholipase A2 | PLA2G2E | 1p36.13 | Q9NZK7 | details | | Group IID secretory phospholipase A2 | PLA2G2D | 1p36.12 | Q9UNK4 | details | | Probable phospholipid-transporting ATPase IG | ATP11C | | Q8NB49 | details | | Probable phospholipid-transporting ATPase IH | ATP11A | 13q34 | P98196 | details | | Probable phospholipid-transporting ATPase VA | ATP10A | 15q11.2 | O60312 | details | | Probable phospholipid-transporting ATPase IC | ATP8B1 | 18q21-q22|18q21.31 | O43520 | details | | Probable phospholipid-transporting ATPase IIA | ATP9A | 20q13.2 | O75110 | details | | Probable phospholipid-transporting ATPase VD | ATP10D | 4p12 | Q9P241 | details | | Probable phospholipid-transporting ATPase IB | ATP8A2 | 13q12 | Q9NTI2 | details | | Probable phospholipid-transporting ATPase IA | ATP8A1 | 4p13 | Q9Y2Q0 | details | | Probable phospholipid-transporting ATPase IM | ATP8B4 | 15q21.2 | Q8TF62 | details | | Probable phospholipid-transporting ATPase IF | ATP11B | 3q27 | Q9Y2G3 | details | | Probable phospholipid-transporting ATPase IK | ATP8B3 | 19p13.3 | O60423 | details | | Choline/ethanolaminephosphotransferase 1 | CEPT1 | 1p13.3 | Q9Y6K0 | details | | Probable phospholipid-transporting ATPase VB | ATP10B | 5q34 | O94823 | details | | Probable phospholipid-transporting ATPase ID | ATP8B2 | 1q21.3 | P98198 | details | | N-acyl-phosphatidylethanolamine-hydrolyzing phospholipase D | NAPEPLD | 7q22.1 | Q6IQ20 | details | | Ethanolaminephosphotransferase 1 | EPT1 | 2p23.3 | Q9C0D9 | details |
|
|---|
| Receptors | |
|---|
| Transcriptional Factors | |
|---|
| Concentrations Data |
|---|
| Not Available |
|---|
| External Links |
|---|
| HMDB ID | HMDB0008932 |
|---|
| DrugBank ID | Not Available |
|---|
| Phenol Explorer Compound ID | Not Available |
|---|
| FoodDB ID | FDB026122 |
|---|
| KNApSAcK ID | Not Available |
|---|
| Chemspider ID | Not Available |
|---|
| KEGG Compound ID | Not Available |
|---|
| BioCyc ID | Not Available |
|---|
| BiGG ID | Not Available |
|---|
| Wikipedia Link | Not Available |
|---|
| METLIN ID | Not Available |
|---|
| PubChem Compound | 9546969 |
|---|
| PDB ID | Not Available |
|---|
| ChEBI ID | 134078 |
|---|
| References |
|---|
| General References | Not Available |
|---|