Showing Compound Card for 2-Oxo-3-phenylpropanoic acid (Mixture oxo and keto) (CDB005145)
| Record Information | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Version | 1.0 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Created at | 2020-04-17 19:13:52 UTC | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Updated at | 2020-12-07 19:11:40 UTC | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CannabisDB ID | CDB005145 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Secondary Accession Numbers | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Cannabis Compound Identification | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Common Name | 2-Oxo-3-phenylpropanoic acid (Mixture oxo and keto) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | Phenylpyruvic acid, also known as keto-phenylpyruvate or a-ketohydrocinnamate, belongs to the class of organic compounds known as phenylpyruvic acid derivatives. Phenylpyruvic acid derivatives are compounds containing a phenylpyruvic acid moiety, which consists of a phenyl group substituted at the second position by an pyruvic acid. Phenylpyruvic acid is an extremely weak basic (essentially neutral) compound (based on its pKa). Phenylpyruvic acid exists in all living species, ranging from bacteria to humans. Within humans, phenylpyruvic acid participates in a number of enzymatic reactions. In particular, phenylpyruvic acid and L-glutamic acid can be biosynthesized from L-phenylalanine and oxoglutaric acid; which is mediated by the enzyme aspartate aminotransferase, cytoplasmic. In addition, phenylpyruvic acid can be biosynthesized from L-phenylalanine through the action of the enzyme L-amino-acid oxidase. In humans, phenylpyruvic acid is involved in the metabolic disorder called tyrosinemia type 3 (tyro3). Phenylpyruvic acid is a potentially toxic compound. Phenylpyruvic acid, with regard to humans, has been found to be associated with the diseases such as primary biliary cirrhosis; phenylpyruvic acid has also been linked to the inborn metabolic disorder phenylketonuria. High levels of phenylpyruvic acid can be found in the urine of individuals with phenylketonuria (PKU), an inborn error of metabolism. Individuals with PKU tend to excrete large quantities of phenylpyruvate, phenylacetate and phenyllactate, along with phenylalanine, in their urine. PKU is due to lack of the enzyme phenylalanine hydroxylase (PAH), so that phenylalanine is converted not to tyrosine but to phenylpyruvic acid. Metabolites of this transamination reaction include phenylacetate, Phenylpyruvic acid and phenethylamine. Thus, excessive levels of phenylalanine significantly decrease the levels of other LNAAs in the brain. The neural-development effects of PKU are primarily due to the disruption of neurotransmitter synthesis. A "musty or mousy" odor of skin, hair, sweat and urine (due to phenylacetate accumulation); and a tendency to hypopigmentation and eczema are also observed. 2-Oxo-3-phenylpropanoic acid (Mixture oxo and keto) is expected to be in Cannabis as all living plants are known to produce and metabolize it. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Structure | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Synonyms |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Formula | C9H8O3 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Average Molecular Weight | 164.16 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Monoisotopic Molecular Weight | 164.0473 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| IUPAC Name | 2-oxo-3-phenylpropanoic acid | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Traditional Name | phenylpyruvic acid | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| CAS Registry Number | 156-06-9 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| SMILES | OC(=O)C(=O)CC1=CC=CC=C1 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Identifier | InChI=1S/C9H8O3/c10-8(9(11)12)6-7-4-2-1-3-5-7/h1-5H,6H2,(H,11,12) | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| InChI Key | BTNMPGBKDVTSJY-UHFFFAOYSA-N | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemical Taxonomy | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Description | Belongs to the class of organic compounds known as phenylpyruvic acid derivatives. Phenylpyruvic acid derivatives are compounds containing a phenylpyruvic acid moiety, which consists of a phenyl group substituted at the second position by an pyruvic acid. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Kingdom | Organic compounds | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Super Class | Benzenoids | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Class | Benzene and substituted derivatives | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Sub Class | Phenylpyruvic acid derivatives | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Direct Parent | Phenylpyruvic acid derivatives | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Alternative Parents | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Substituents |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Molecular Framework | Aromatic homomonocyclic compounds | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Descriptors |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Ontology | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physiological effect | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Disposition | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Role | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Physical Properties | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| State | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Experimental Properties |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Predicted Properties |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Spectra | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| EI-MS/GC-MS |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| MS/MS |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| NMR |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
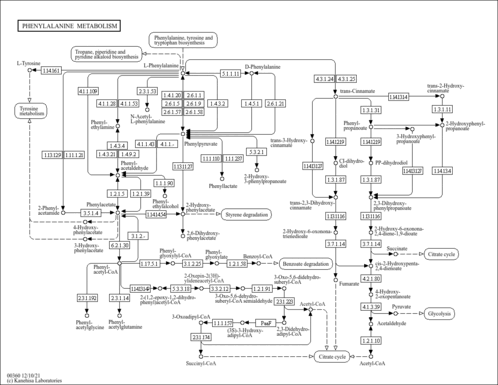
| Pathways | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Pathways | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Protein Targets | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Enzymes |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Transporters | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Metal Bindings |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Receptors |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Transcriptional Factors | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Concentrations Data | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Not Available | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| External Links | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| HMDB ID | HMDB0000205 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| DrugBank ID | DB03884 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Phenol Explorer Compound ID | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| FoodDB ID | FDB008272 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KNApSAcK ID | C00000751 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Chemspider ID | 972 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| KEGG Compound ID | C00166 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| BioCyc ID | PHENYL-PYRUVATE | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| BiGG ID | 34111 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikipedia Link | Phenylpyruvic_acid | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| METLIN ID | 328 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PubChem Compound | 997 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PDB ID | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ChEBI ID | 30851 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| References | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| General References | Not Available | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Enzymes
- General function:
- Involved in cell surface binding
- Specific function:
- Pro-inflammatory cytokine. Involved in the innate immune response to bacterial pathogens. The expression of MIF at sites of inflammation suggests a role as mediator in regulating the function of macrophages in host defense. Counteracts the anti-inflammatory activity of glucocorticoids. Has phenylpyruvate tautomerase and dopachrome tautomerase activity (in vitro), but the physiological substrate is not known. It is not clear whether the tautomerase activity has any physiological relevance, and whether it is important for cytokine activity.
- Gene Name:
- MIF
- Uniprot ID:
- P14174
- Molecular weight:
- 12476.19
- General function:
- Involved in 1-aminocyclopropane-1-carboxylate synthase activity
- Specific function:
- Transaminase involved in tyrosine breakdown. Converts tyrosine to p-hydroxyphenylpyruvate. Can catalyze the reverse reaction, using glutamic acid, with 2-oxoglutarate as cosubstrate (in vitro). Has much lower affinity and transaminase activity towards phenylalanine.
- Gene Name:
- TAT
- Uniprot ID:
- P17735
- Molecular weight:
- 50398.895
- General function:
- Involved in transferase activity, transferring nitrogenous groups
- Specific function:
- Plays a key role in amino acid metabolism (By similarity).
- Gene Name:
- GOT1
- Uniprot ID:
- P17174
- Molecular weight:
- 46247.14
- General function:
- Involved in transferase activity, transferring nitrogenous groups
- Specific function:
- Catalyzes the irreversible transamination of the L-tryptophan metabolite L-kynurenine to form kynurenic acid (KA). Plays a key role in amino acid metabolism. Important for metabolite exchange between mitochondria and cytosol. Facilitates cellular uptake of long-chain free fatty acids.
- Gene Name:
- GOT2
- Uniprot ID:
- P00505
- Molecular weight:
- 47517.285
- General function:
- Involved in 1-aminocyclopropane-1-carboxylate synthase activity
- Specific function:
- Catalyzes the irreversible transamination of the L-tryptophan metabolite L-kynurenine to form kynurenic acid (KA). Metabolizes the cysteine conjugates of certain halogenated alkenes and alkanes to form reactive metabolites. Catalyzes the beta-elimination of S-conjugates and Se-conjugates of L-(seleno)cysteine, resulting in the cleavage of the C-S or C-Se bond.
- Gene Name:
- CCBL1
- Uniprot ID:
- Q16773
- Molecular weight:
- 47874.765
- General function:
- Involved in transferase activity, transferring hexosyl groups
- Specific function:
- UDPGT is of major importance in the conjugation and subsequent elimination of potentially toxic xenobiotics and endogenous compounds. This isoform glucuronidates bilirubin IX-alpha to form both the IX-alpha-C8 and IX-alpha-C12 monoconjugates and diconjugate. Is also able to catalyze the glucuronidation of 17beta-estradiol, 17alpha-ethinylestradiol, 1-hydroxypyrene, 4-methylumbelliferone, 1-naphthol, paranitrophenol, scopoletin, and umbelliferone.
- Gene Name:
- UGT1A1
- Uniprot ID:
- P22309
- Molecular weight:
- 59590.91
- General function:
- Involved in 4-hydroxyphenylpyruvate dioxygenase activity
- Specific function:
- Key enzyme in the degradation of tyrosine.
- Gene Name:
- HPD
- Uniprot ID:
- P32754
- Molecular weight:
- 40497.105
- General function:
- Involved in oxidoreductase activity
- Specific function:
- Lysosomal L-amino-acid oxidase with highest specific activity with phenylalanine. May play a role in lysosomal antigen processing and presentation (By similarity).
- Gene Name:
- IL4I1
- Uniprot ID:
- Q96RQ9
- Molecular weight:
- 65327.26