| Record Information |
|---|
| Version | 1.0 |
|---|
| Created at | 2020-04-17 19:08:37 UTC |
|---|
| Updated at | 2020-11-18 16:39:20 UTC |
|---|
| CannabisDB ID | CDB005093 |
|---|
| Secondary Accession Numbers | Not Available |
|---|
| Cannabis Compound Identification |
|---|
| Common Name | FADH |
|---|
| Description | FADH, also known as 1,5-dihydro-fad or FADH2, belongs to the class of organic compounds known as flavin nucleotides. These are nucleotides containing a flavin moiety. Flavin is a compound that contains the tricyclic isoalloxazine ring system, which bears 2 oxo groups at the 2- and 4-positions. FADH is a strong basic compound (based on its pKa). FADH exists in all living species, ranging from bacteria to humans. In humans, FADH is involved in lps and citrate signaling and inflammation. Outside of the human body, FADH has been detected, but not quantified in, several different foods, such as thistles, white lupines, grass pea, deerberries, and passion fruits. This could make FADH a potential biomarker for the consumption of these foods. FADH is expected to be in Cannabis as all living plants are known to produce and metabolize it. |
|---|
| Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| 1,5-Dihydro-fad | ChEBI | | DIHYDROFLAVINE-adenine dinucleotide | ChEBI | | Flavin adenine dinucleotide (reduced) | ChEBI | | 1,5-Dihydro-p-5-ester with adenosine | HMDB | | 1,5-Dihydro-riboflavin 5'-(trihydrogen diphosphate) p'->5'-ester with adenosine | HMDB | | Adenosine 5'-(trihydrogen pyrophosphate), 5'-5'-ester with 5,10-dihydro-7,8-dimethyl-10-(D-ribo-2,3,4,5-tetrahydroxypentyl)alloxazine | HMDB | | Adenosine 5'-(trihydrogen pyrophosphate), 5'->5'-ester with 5,10-dihydro-7,8-dimethyl-10-(D-ribo-2,3,4,5-tetrahydroxypentyl)alloxazine | HMDB | | Adenosine 5'-{3-[D-ribo-5-(7,8-dimethyl-2,4-dioxo-1,2,3,4,5,10-tetrahydrobenzo[g]pteridin-10-yl)-2,3,4-trihydroxypentyl] dihydrogen diphosphate} | HMDB | | Adenosine 5-(trihydrogen pyrophosphate) | HMDB | | Adenosine pyrophosphate 5'-5'-ester with 5,10-dihydro-7,8-dimethyl-10-(D-ribo-2,3,4,5-tetrahydroxypentyl)alloxazine | HMDB | | Adenosine pyrophosphate, 5'-5'-ester with 5,10-dihydro-7,8-dimethyl-10-(D-ribo-2,3,4,5-tetrahydroxypentyl)alloxazine | HMDB | | Adenosine pyrophosphate, 5'->5'-ester with 5,10-dihydro-7,8-dimethyl-10-(D-ribo-2,3,4,5-tetrahydroxypentyl)alloxazine | HMDB | | Benzo[GR]pteridine riboflavin 5'-(trihydrogen diphosphate) deriv | HMDB | | Benzo[g]pteridine riboflavin 5'-(trihydrogen diphosphate) deriv | HMDB | | Dihydro-fad | HMDB | | FADH2 | HMDB | | FDA | HMDB | | Flavin adenine dinucleotide reduced | HMDB | | Reduced flavine adenine dinucleotide | HMDB |
|
|---|
| Chemical Formula | C27H35N9O15P2 |
|---|
| Average Molecular Weight | 787.57 |
|---|
| Monoisotopic Molecular Weight | 787.1728 |
|---|
| IUPAC Name | {[(2R,3S,4R,5R)-5-(6-amino-9H-purin-9-yl)-3,4-dihydroxyoxolan-2-yl]methoxy}[({[(2R,3S,4S)-5-{7,8-dimethyl-2,4-dioxo-1H,2H,3H,4H,5H,10H-benzo[g]pteridin-10-yl}-2,3,4-trihydroxypentyl]oxy}(hydroxy)phosphoryl)oxy]phosphinic acid |
|---|
| Traditional Name | fadh(.) |
|---|
| CAS Registry Number | 1910-41-4 |
|---|
| SMILES | CC1=CC2=C(C=C1C)N(C[C@H](O)[C@H](O)[C@H](O)COP(O)(=O)OP(O)(=O)OC[C@H]1O[C@H]([C@H](O)[C@@H]1O)N1C=NC3=C1N=CN=C3N)C1=C(N2)C(=O)NC(=O)N1 |
|---|
| InChI Identifier | InChI=1S/C27H35N9O15P2/c1-10-3-12-13(4-11(10)2)35(24-18(32-12)25(42)34-27(43)33-24)5-14(37)19(39)15(38)6-48-52(44,45)51-53(46,47)49-7-16-20(40)21(41)26(50-16)36-9-31-17-22(28)29-8-30-23(17)36/h3-4,8-9,14-16,19-21,26,32,37-41H,5-7H2,1-2H3,(H,44,45)(H,46,47)(H2,28,29,30)(H2,33,34,42,43)/t14-,15+,16+,19-,20+,21+,26+/m0/s1 |
|---|
| InChI Key | YPZRHBJKEMOYQH-UYBVJOGSSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | Belongs to the class of organic compounds known as flavin nucleotides. These are nucleotides containing a flavin moiety. Flavin is a compound that contains the tricyclic isoalloxazine ring system, which bears 2 oxo groups at the 2- and 4-positions. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Nucleosides, nucleotides, and analogues |
|---|
| Class | Flavin nucleotides |
|---|
| Sub Class | Not Available |
|---|
| Direct Parent | Flavin nucleotides |
|---|
| Alternative Parents | |
|---|
| Substituents | - Flavin nucleotide
- Purine ribonucleoside diphosphate
- Purine ribonucleoside monophosphate
- Flavin
- Pentose phosphate
- Pentose-5-phosphate
- Alkyldiarylamine
- Glycosyl compound
- N-glycosyl compound
- 6-aminopurine
- Organic pyrophosphate
- Pentose monosaccharide
- Pteridine
- Monosaccharide phosphate
- Imidazopyrimidine
- Purine
- Aminopyrimidine
- Monoalkyl phosphate
- Pyrimidone
- Imidolactam
- Benzenoid
- Phosphoric acid ester
- Organic phosphoric acid derivative
- N-substituted imidazole
- Monosaccharide
- Pyrimidine
- Alkyl phosphate
- Azole
- Heteroaromatic compound
- Tetrahydrofuran
- Imidazole
- Vinylogous amide
- Urea
- Lactam
- Secondary alcohol
- Secondary amine
- Organoheterocyclic compound
- Azacycle
- Oxacycle
- Polyol
- Primary amine
- Amine
- Alcohol
- Organic nitrogen compound
- Hydrocarbon derivative
- Organic oxide
- Organopnictogen compound
- Organic oxygen compound
- Organooxygen compound
- Organonitrogen compound
- Aromatic heteropolycyclic compound
|
|---|
| Molecular Framework | Aromatic heteropolycyclic compounds |
|---|
| External Descriptors | |
|---|
| Ontology |
|---|
|
| Disposition | Route of exposure: Source: |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Experimental Properties | | Property | Value | Reference |
|---|
| Melting Point | Not Available | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | Not Available | Not Available | | logP | -1.336 | Wikipedia |
|
|---|
| Predicted Properties | [] |
|---|
| Spectra |
|---|
| EI-MS/GC-MS | | Type | Description | Splash Key | View |
|---|
| Predicted GC-MS | FADH, non-derivatized, Predicted GC-MS Spectrum - 70eV, Positive | splash10-0g70-0024960400-5de09e4bdf0187b78529 | Spectrum | | Predicted GC-MS | FADH, TMS_1_1, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | FADH, TMS_1_2, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | FADH, TMS_1_3, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | FADH, TMS_1_4, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | FADH, TMS_1_5, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | FADH, TMS_1_6, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | FADH, TMS_1_7, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | FADH, TMS_1_8, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | FADH, TMS_1_9, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | FADH, TMS_1_10, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum | | Predicted GC-MS | FADH, TMS_1_11, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum |
|
|---|
| MS/MS | | Type | Description | Splash Key | View |
|---|
| MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 10V, Positive (Annotated) | splash10-000i-0001200900-b0740b3d33d50996ccde | 2012-07-24 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 25V, Positive (Annotated) | splash10-000m-0105900000-3abff26d5bb35f8527b8 | 2012-07-24 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 40V, Positive (Annotated) | splash10-000i-0931700000-73f360589a13230eea7a | 2012-07-24 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-000i-0932110400-58a2ec43460cf9df1c4d | 2016-09-12 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-000i-0930000000-15d7e9b22a8286323cc5 | 2016-09-12 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-000i-0960000000-64746c5025e9c53c89b3 | 2016-09-12 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0159-1675510900-39158f1ef2e33daa689e | 2016-09-12 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-001l-4940100000-64cc07487da7b6c45ba2 | 2016-09-12 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0a7i-4900000000-4da9166c58a097f7d7ef | 2016-09-12 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-000i-0104000900-487b1a93da5295026592 | 2021-09-22 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-000i-0964110400-9a24ca36f159cc16d182 | 2021-09-22 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-052u-0693100000-d2fe9957910ae7607fd3 | 2021-09-22 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-000i-0000000900-5c202a395443de89e132 | 2021-09-23 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-005i-9611010700-e3e8cae89c8f03508db4 | 2021-09-23 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-05r0-9735640500-89a663c6bc64f3d8f9f1 | 2021-09-23 | View Spectrum |
|
|---|
| NMR | | Type | Description | | View |
|---|
| 1D NMR | 1H NMR Spectrum (1D, 500 MHz, H2O, experimental) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 100 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 100 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 1000 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 200 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 200 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 300 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 300 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 400 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 500 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 500 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 600 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 600 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 700 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 700 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 800 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 800 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 13C NMR Spectrum (1D, 900 MHz, D2O, predicted) | | Spectrum | | 1D NMR | 1H NMR Spectrum (1D, 900 MHz, D2O, predicted) | | Spectrum | | 2D NMR | [1H, 1H]-TOCSY. Unexported temporarily by An Chi on Oct 15, 2021 until json or nmrML file is generated. 2D NMR Spectrum (experimental) | | Spectrum | | 2D NMR | [1H, 13C]-HSQC NMR Spectrum (2D, 600 MHz, H2O, experimental) | | Spectrum |
|
|---|
| Pathways |
|---|
| Pathways | | Name | SMPDB/Pathwhiz | KEGG | | Citric Acid Cycle | 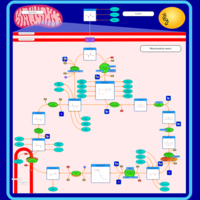 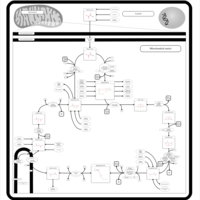  | 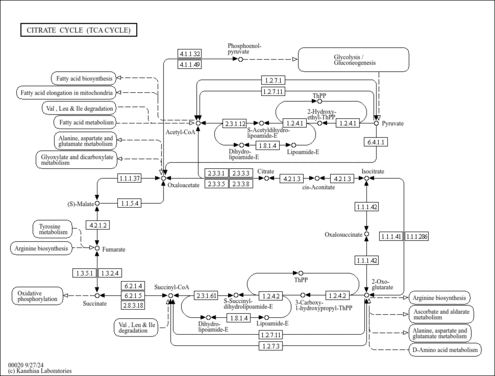 | | Glycerol Phosphate Shuttle | 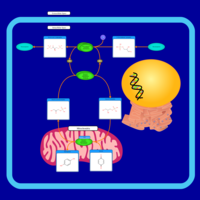 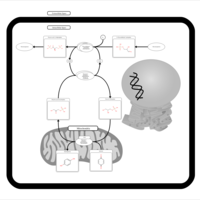  | Not Available | | Congenital lactic acidosis | 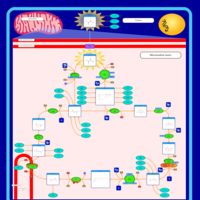 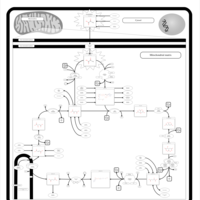 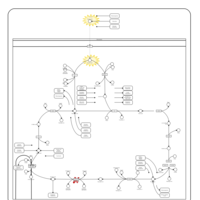 | Not Available | | Fumarase deficiency | 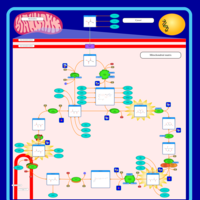 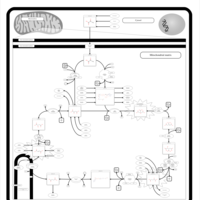 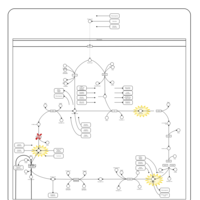 | Not Available | | Mitochondrial complex II deficiency | 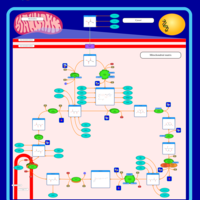 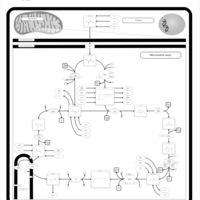 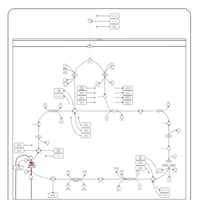 | Not Available |
|
|---|
| Protein Targets |
|---|
| Enzymes | |
| Long-chain specific acyl-CoA dehydrogenase, mitochondrial | ACADL | 2q34 | P28330 | details | | Short-chain specific acyl-CoA dehydrogenase, mitochondrial | ACADS | 12q24.31 | P16219 | details | | Medium-chain specific acyl-CoA dehydrogenase, mitochondrial | ACADM | 1p31 | P11310 | details | | Peroxisomal acyl-coenzyme A oxidase 1 | ACOX1 | 17q25.1 | Q15067 | details | | Heme oxygenase 2 | HMOX2 | 16p13.3 | P30519 | details | | Isovaleryl-CoA dehydrogenase, mitochondrial | IVD | 15q14-q15 | P26440 | details | | Peroxisomal acyl-coenzyme A oxidase 3 | ACOX3 | 4p15.3 | O15254 | details | | Glycerol-3-phosphate dehydrogenase, mitochondrial | GPD2 | 2q24.1 | P43304 | details | | Heme oxygenase 1 | HMOX1 | 22q13.1 | P09601 | details | | Glutaryl-CoA dehydrogenase, mitochondrial | GCDH | 19p13.2 | Q92947 | details | | Flavin reductase (NADPH) | BLVRB | 19q13.1-q13.2 | P30043 | details | | Short/branched chain specific acyl-CoA dehydrogenase, mitochondrial | ACADSB | 10q26.13 | P45954 | details | | L-2-hydroxyglutarate dehydrogenase, mitochondrial | L2HGDH | 14q21.3 | Q9H9P8 | details | | Very long-chain specific acyl-CoA dehydrogenase, mitochondrial | ACADVL | 17p13.1 | P49748 | details |
|
|---|
| Transporters | Not Available |
|---|
| Metal Bindings | |
|---|
| Receptors | Not Available |
|---|
| Transcriptional Factors | |
|---|
| Concentrations Data |
|---|
| Not Available |
|---|
| External Links |
|---|
| HMDB ID | HMDB0001197 |
|---|
| DrugBank ID | Not Available |
|---|
| Phenol Explorer Compound ID | Not Available |
|---|
| FoodDB ID | FDB030853 |
|---|
| KNApSAcK ID | Not Available |
|---|
| Chemspider ID | 393487 |
|---|
| KEGG Compound ID | C01352 |
|---|
| BioCyc ID | FADH2 |
|---|
| BiGG ID | 132077 |
|---|
| Wikipedia Link | Flavin adenine dinucleotide |
|---|
| METLIN ID | 6073 |
|---|
| PubChem Compound | 446013 |
|---|
| PDB ID | Not Available |
|---|
| ChEBI ID | 17877 |
|---|
| References |
|---|
| General References | Not Available |
|---|
