| Identification |
|---|
| HMDB Protein ID
| CDBP00956 |
| Secondary Accession Numbers
| Not Available |
| Name
| Cytochrome P450 3A4 |
| Description
| Not Available |
| Synonyms
|
- Albendazole monooxygenase
- Albendazole sulfoxidase
- CYPIIIA3
- CYPIIIA4
- Cytochrome P450 3A3
- Cytochrome P450 HLp
- Cytochrome P450 NF-25
- Cytochrome P450-PCN1
- Nifedipine oxidase
- Quinine 3-monooxygenase
- Taurochenodeoxycholate 6-alpha-hydroxylase
- 1,8-cineole 2-exo-monooxygenase
|
| Gene Name
| CYP3A4 |
| Protein Type
| Metal Binding |
| Biological Properties |
|---|
| General Function
| Involved in monooxygenase activity |
| Specific Function
| Cytochromes P450 are a group of heme-thiolate monooxygenases. In liver microsomes, this enzyme is involved in an NADPH-dependent electron transport pathway. It performs a variety of oxidation reactions (e.g. caffeine 8-oxidation, omeprazole sulphoxidation, midazolam 1'-hydroxylation and midazolam 4-hydroxylation) of structurally unrelated compounds, including steroids, fatty acids, and xenobiotics. Acts as a 1,8-cineole 2-exo-monooxygenase. The enzyme also hydroxylates etoposide.
|
| GO Classification
|
| Biological Process |
| alkaloid catabolic process |
| androgen metabolic process |
| vitamin D metabolic process |
| oxidative demethylation |
| exogenous drug catabolic process |
| heterocycle metabolic process |
| monoterpenoid metabolic process |
| steroid catabolic process |
| xenobiotic metabolic process |
| Cellular Component |
| endoplasmic reticulum membrane |
| cell surface |
| integral to membrane |
| Function |
| oxidoreductase activity |
| ion binding |
| cation binding |
| metal ion binding |
| binding |
| catalytic activity |
| transition metal ion binding |
| electron carrier activity |
| iron ion binding |
| monooxygenase activity |
| heme binding |
| oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced flavin or flavoprotein as one donor, and incorporation of one atom of oxygen |
| Molecular Function |
| electron carrier activity |
| oxygen binding |
| vitamin D 24-hydroxylase activity |
| caffeine oxidase activity |
| oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced flavin or flavoprotein as one donor, and incorporation of one atom of oxygen |
| iron ion binding |
| vitamin D3 25-hydroxylase activity |
| heme binding |
| albendazole monooxygenase activity |
| quinine 3-monooxygenase activity |
| taurochenodeoxycholate 6alpha-hydroxylase activity |
| testosterone 6-beta-hydroxylase activity |
| steroid binding |
| Process |
| metabolic process |
| oxidation reduction |
|
| Cellular Location
|
- Endoplasmic reticulum membrane
- Single-pass membrane protein
- Single-pass membrane protein
- Microsome membrane
|
| Pathways
|
| Name | SMPDB/Pathwhiz | KEGG | | Steroid hormone biosynthesis | Not Available | 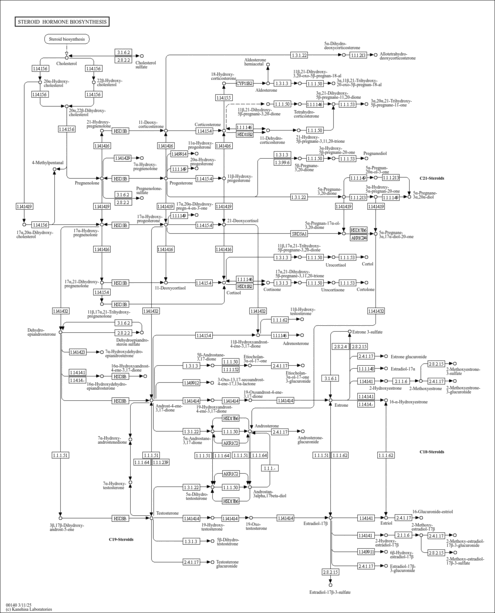 | | Linoleic acid metabolism | Not Available | 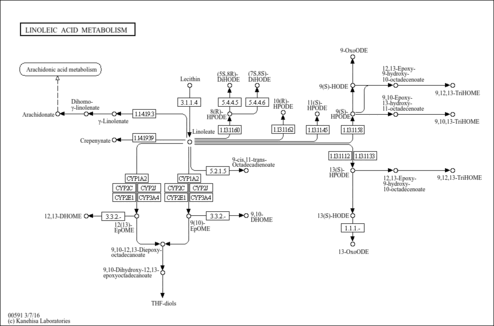 | | Metabolism of xenobiotics by cytochrome P450 | Not Available | 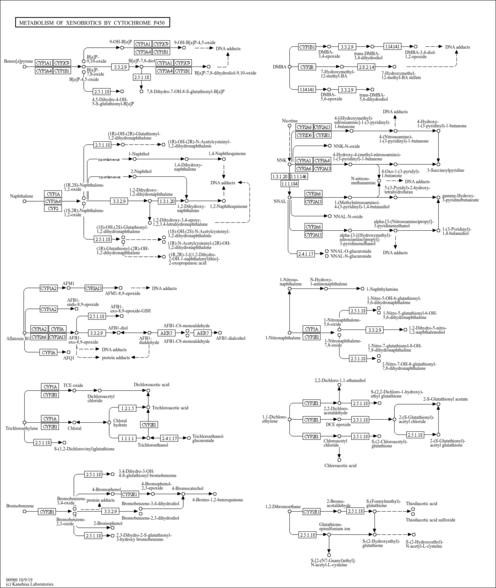 | | Drug metabolism - cytochrome P450 | Not Available | 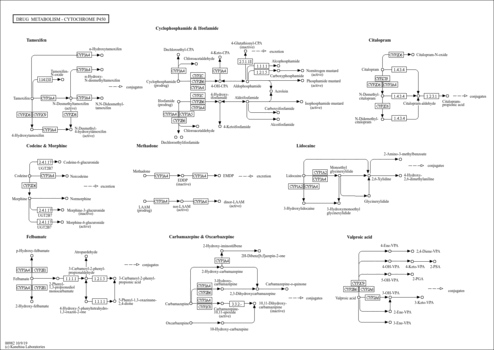 | | Drug metabolism - other enzymes | Not Available | 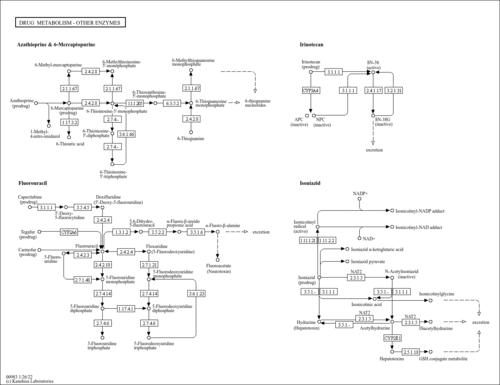 |
|
| Gene Properties |
|---|
| Chromosome Location
| 7 |
| Locus
| 7q21.1 |
| SNPs
| CYP3A4 |
| Gene Sequence
|
>1512 bp
ATGGCTCTCATCCCAGACTTGGCCATGGAAACCCGGCTTCTCCTGGCTGTCAGCCTGGTG
CTCCTCTATCTATATGGAACTCATTCACATGGACTTTTTAAGAAGCTTGGAATTCCAGGG
CCCACACCTCTGCCTTTTTTGGGAAATATTTTGTCCTACCATAAGGGCTTTTGTATGTTT
GACATGGAATGTCATAAAAAGTATGGAAAAGTGTGGGGCTTTTATGATGGTCAACAGCCT
GTGCTGGCTATCACAGATCCTGACATGATCAAAACAGTGCTAGTGAAAGAATGTTATTCT
GTCTTCACAAACCGGAGGCCTTTTGGTCCAGTGGGATTTATGAAAAGTGCCATCTCTATA
GCTGAGGATGAAGAATGGAAGAGATTACGATCATTGCTGTCTCCAACCTTCACCAGTGGA
AAACTCAAGGAGATGGTCCCTATCATTGCCCAGTATGGAGATGTGTTGGTGAGAAATCTG
AGGCGGGAAGCAGAGACAGGCAAGCCTGTCACCTTGAAAGACGTCTTTGGGGCCTACAGC
ATGGATGTGATCACTAGCACATCATTTGGAGTGAACATCGACTCTCTCAACAATCCACAA
GACCCCTTTGTGGAAAACACCAAGAAGCTTTTAAGATTTGATTTTTTGGATCCATTCTTT
CTCTCAATAACAGTCTTTCCATTCCTCATCCCAATTCTTGAAGTATTAAATATCTGTGTG
TTTCCAAGAGAAGTTACAAATTTTTTAAGAAAATCTGTAAAAAGGATGAAAGAAAGTCGC
CTCGAAGATACACAAAAGCACCGAGTGGATTTCCTTCAGCTGATGATTGACTCTCAGAAT
TCAAAAGAAACTGAGTCCCACAAAGCTCTGTCCGATCTGGAGCTCGTGGCCCAATCAATT
ATCTTTATTTTTGCTGGCTGTGAAACCACGAGCAGTGTTCTCTCCTTCATTATGTATGAA
CTGGCCACTCACCCTGATGTCCAGCAGAAACTGCAGGAGGAAATTGATGCAGTTTTACCC
AATAAGGCACCACCCACCTATGATACTGTGCTACAGATGGAGTATCTTGACATGGTGGTG
AATGAAACGCTCAGATTATTCCCAATTGCTATGAGACTTGAGAGGGTCTGCAAAAAAGAT
GTTGAGATCAATGGGATGTTCATTCCCAAAGGGGTGGTGGTGATGATTCCAAGCTATGCT
CTTCACCGTGACCCAAAGTACTGGACAGAGCCTGAGAAGTTCCTCCCTGAAAGATTCAGC
AAGAAGAACAAGGACAACATAGATCCTTACATATACACACCCTTTGGAAGTGGACCCAGA
AACTGCATTGGCATGAGGTTTGCTCTCATGAACATGAAACTTGCTCTAATCAGAGTCCTT
CAGAACTTCTCCTTCAAACCTTGTAAAGAAACACAGATCCCCCTGAAATTAAGCTTAGGA
GGACTTCTTCAACCAGAAAAACCCGTTGTTCTAAAGGTTGAGTCAAGGGATGGCACCGTA
AGTGGAGCCTGA
|
| Protein Properties |
|---|
| Number of Residues
| 503 |
| Molecular Weight
| 57255.585 |
| Theoretical pI
| 8.097 |
| Pfam Domain Function
|
|
| Signals
|
Not Available
|
|
Transmembrane Regions
|
Not Available
|
| Protein Sequence
|
>Cytochrome P450 3A4
MALIPDLAMETWLLLAVSLVLLYLYGTHSHGLFKKLGIPGPTPLPFLGNILSYHKGFCMF
DMECHKKYGKVWGFYDGQQPVLAITDPDMIKTVLVKECYSVFTNRRPFGPVGFMKSAISI
AEDEEWKRLRSLLSPTFTSGKLKEMVPIIAQYGDVLVRNLRREAETGKPVTLKDVFGAYS
MDVITSTSFGVNIDSLNNPQDPFVENTKKLLRFDFLDPFFLSITVFPFLIPILEVLNICV
FPREVTNFLRKSVKRMKESRLEDTQKHRVDFLQLMIDSQNSKETESHKALSDLELVAQSI
IFIFAGYETTSSVLSFIMYELATHPDVQQKLQEEIDAVLPNKAPPTYDTVLQMEYLDMVV
NETLRLFPIAMRLERVCKKDVEINGMFIPKGVVVMIPSYALHRDPKYWTEPEKFLPERFS
KKNKDNIDPYIYTPFGSGPRNCIGMRFALMNMKLALIRVLQNFSFKPCKETQIPLKLSLG
GLLQPEKPVVLKVESRDGTVSGA
|
| External Links |
|---|
| GenBank ID Protein
| 6470135 |
| UniProtKB/Swiss-Prot ID
| P08684 |
| UniProtKB/Swiss-Prot Entry Name
| CP3A4_HUMAN |
| PDB IDs
|
|
| GenBank Gene ID
| AF182273 |
| GeneCard ID
| CYP3A4 |
| GenAtlas ID
| CYP3A4 |
| HGNC ID
| HGNC:2637 |
| References |
|---|
| General References
| Not Available |




