| Identification |
|---|
| HMDB Protein ID
| CDBP03920 |
| Secondary Accession Numbers
| Not Available |
| Name
| m7GpppN-mRNA hydrolase |
| Description
| Not Available |
| Synonyms
|
- Nucleoside diphosphate-linked moiety X motif 20
- Nudix motif 20
- hDpc
- mRNA-decapping enzyme 2
|
| Gene Name
| DCP2 |
| Protein Type
| Enzyme |
| Biological Properties |
|---|
| General Function
| Involved in RNA binding |
| Specific Function
| Necessary for the degradation of mRNAs, both in normal mRNA turnover and in nonsense-mediated mRNA decay. Plays a role in replication-dependent histone mRNA degradation. Removes the 7-methyl guanine cap structure from mRNA molecules, yielding a 5'-phosphorylated mRNA fragment and 7m-GDP. Has higher activity towards mRNAs that lack a poly(A) tail. Has no activity towards a cap structure lacking a RNA moiety.
|
| GO Classification
|
| Biological Process |
| gene expression |
| histone mRNA catabolic process |
| nuclear-transcribed mRNA catabolic process, nonsense-mediated decay |
| exonucleolytic nuclear-transcribed mRNA catabolic process involved in deadenylation-dependent decay |
| Cellular Component |
| cytosol |
| cytoplasmic mRNA processing body |
| RNA-induced silencing complex |
| nucleus |
| Function |
| rna binding |
| manganese ion binding |
| ion binding |
| cation binding |
| metal ion binding |
| binding |
| catalytic activity |
| hydrolase activity |
| transition metal ion binding |
| nucleic acid binding |
| Molecular Function |
| exoribonuclease activity, producing 5'-phosphomonoesters |
| m7G(5')pppN diphosphatase activity |
| manganese ion binding |
| RNA binding |
|
| Cellular Location
|
- Nucleus
- Cytoplasm
- P-body
|
| Pathways
|
| Name | SMPDB/Pathwhiz | KEGG | | RNA degradation | Not Available | 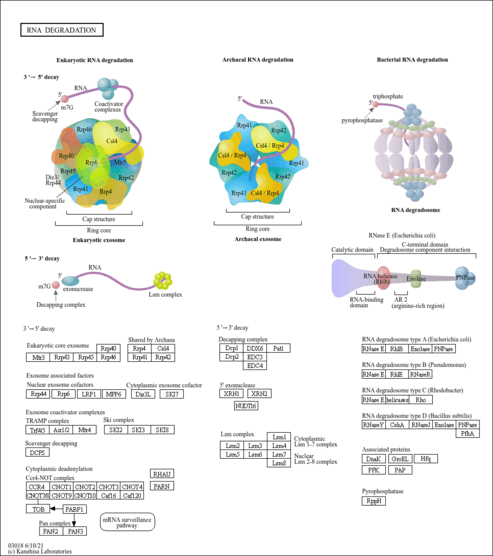 |
|
| Gene Properties |
|---|
| Chromosome Location
| 5 |
| Locus
| 5q22.2 |
| SNPs
| DCP2 |
| Gene Sequence
|
>1263 bp
ATGGAGACCAAACGGGTGGAGATTCCCGGCAGCGTCCTGGACGATCTCTGCAGCCGATTT
ATTTTGCATATTCCCAGCGAGGAAAGAGACAATGCAATCCGAGTGTGTTTTCAGATTGAA
CTTGCCCATTGGTTTTACTTGGATTTCTACATGCAGAACACACCAGGATTACCTCAGTGT
GGGATAAGAGACTTTGCTAAAGCTGTCTTCAGTCATTGTCCGTTTTTGCTGCCTCAAGGT
GAAGATGTGGAAAAAGTTTTGGATGAATGGAAGGAATATAAAATGGGAGTACCAACATAT
GGTGCAATTATTCTTGATGAGACACTTGAAAATGTACTACTAGTTCAGGGGTACCTAGCA
AAATCAGGCTGGGGATTTCCAAAAGGAAAAGTAAATAAAGAAGAAGCTCCTCATGATTGT
GCTGCTAGAGAGGTCTTTGAAGAAACTGGTTTTGATATCAAAGACTATATTTGTAAGGAT
GATTACATTGAACTTCGAATCAATGACCAGCTTGCTCGTTTGTACATCATTCCAGGAATT
CCAAAAGACACAAAATTTAACCCAAAAACTAGAAGAGAAATTCGGAACATTGAGTGGTTC
TCTATTGAGAAATTGCCTTGTCATAGAAATGATATGACCCCCAAATCCAAACTTGGTTTG
GCACCTAACAAATTTTTTATGGCCATTCCCTTTATCAGACCATTAAGGGACTGGCTTTCT
CGAAGATTTGGCGATTCCTCAGACAGTGACAATGGATTTTCCTCAACTGGTAGCACGCCG
GCTAAACCCACTGTGGAAAAATTGAGTCGAACCAAATTCCGCCACAGTCAGCAGTTATTT
CCTGACGGTTCTCCTGGTGACCAGTGGGTAAAGCACAGGCAACCACTGCAGCAAAAGCCA
TATAATAATCATTCTGAAATGTCTGACCTTTTAAAAGGAAAGAATCAAAGTATGAGGGGA
AATGGCAGAAAACAGTATCAAGATTCACCTAATCAAAAGAAAAGAACAAATGGGCTTCAG
CCAGCAAAGCAGCAGAATTCTTTGATGAAGTGTGAAAAGAAACTTCATCCACGGAAACTT
CAGGATAATTTTGAAACAGATGCTGTATATGACTTGCCTAGCTCCAGTGAAGACCAGTTG
CTAGAACATGCTGAGGGACAGCCCGTGGCATGTAATGGACATTGCAAGTTCCCCTTTTCA
TCCAGAGCCTTTTTGAGTTTCAAGTTTGACCATAATGCTATAATGAAAATCTTGGACCTT
TGA
|
| Protein Properties |
|---|
| Number of Residues
| 420 |
| Molecular Weight
| 44408.25 |
| Theoretical pI
| 6.936 |
| Pfam Domain Function
|
|
| Signals
|
Not Available
|
|
Transmembrane Regions
|
Not Available
|
| Protein Sequence
|
>mRNA-decapping enzyme 2
METKRVEIPGSVLDDFCSRFILHIPSEERDNAIRVCFQIELAHWFYLDFYMQNTPGLPQC
GIRDFAKAVFSHCPFLLPQGEDVEKVLDEWKEYKMGVPTYGAIILDETLENVLLVQGYLA
KSGWGFPKGKVNKEEAPHDCAAREVFEETGFDIKDYICKDDYIELRINDQLARLYIIPGI
PKDTKFNPKTRREIRNIEWFSIEKLPCHRNDMTPKSKLGLAPNKFFMAIPFIRPLRDWLS
RRFGDSSDSDNGFSSTGSTPAKPTVEKLSRTKFRHSQQLFPDGSPGDQWVKHRQPLQQKP
YNNHSEMSDLLKGKNQSMRGNGRKQYQDSPNQKKRTNGLQPAKQQNSLMKCEKKLHPRKL
QDNFETDAVYDLPSSSEDQLLEHAEGQPVACNGHCKFPFSSRAFLSFKFDHNAIMKILDL
|
| External Links |
|---|
| GenBank ID Protein
| 31542498 |
| UniProtKB/Swiss-Prot ID
| Q8IU60 |
| UniProtKB/Swiss-Prot Entry Name
| DCP2_HUMAN |
| PDB IDs
|
Not Available |
| GenBank Gene ID
| NM_152624.4 |
| GeneCard ID
| DCP2 |
| GenAtlas ID
| DCP2 |
| HGNC ID
| HGNC:24452 |
| References |
|---|
| General References
| Not Available |
