| Identification |
|---|
| HMDB Protein ID
| CDBP00808 |
| Secondary Accession Numbers
| Not Available |
| Name
| Inositol monophosphatase 1 |
| Description
| Not Available |
| Synonyms
|
- IMP 1
- IMPase 1
- Inositol-1(or 4)-monophosphatase 1
- Lithium-sensitive myo-inositol monophosphatase A1
|
| Gene Name
| IMPA1 |
| Protein Type
| Enzyme |
| Biological Properties |
|---|
| General Function
| Involved in inositol or phosphatidylinositol phosphatase activity |
| Specific Function
| Responsible for the provision of inositol required for synthesis of phosphatidylinositol and polyphosphoinositides and has been implicated as the pharmacological target for lithium action in brain. Can use myo-inositol monophosphates, myo-inositol 1,3-diphosphate, myo-inositol 1,4-diphosphate, scyllo-inositol-phosphate, glucose-1-phosphate, glucose-6-phosphate, fructose-1-phosphate, beta-glycerophosphate, and 2'-AMP as substrates.
|
| GO Classification
|
| Biological Process |
| response to lithium ion |
| signal transduction |
| phosphatidylinositol biosynthetic process |
| inositol biosynthetic process |
| inositol phosphate dephosphorylation |
| phosphatidylinositol phosphorylation |
| Cellular Component |
| mitochondrion |
| nucleus |
| neuronal cell body |
| axon |
| Function |
| hydrolase activity, acting on ester bonds |
| catalytic activity |
| hydrolase activity |
| phosphoric ester hydrolase activity |
| phosphatase activity |
| inositol or phosphatidylinositol phosphatase activity |
| Molecular Function |
| metal ion binding |
| inositol monophosphate 1-phosphatase activity |
| inositol monophosphate 3-phosphatase activity |
| inositol monophosphate 4-phosphatase activity |
|
| Cellular Location
|
- Cytoplasm
|
| Pathways
|
| Name | SMPDB/Pathwhiz | KEGG | | myo-inositol biosynthesis | Not Available | Not Available | | Phosphatidylinositol signaling system | Not Available | 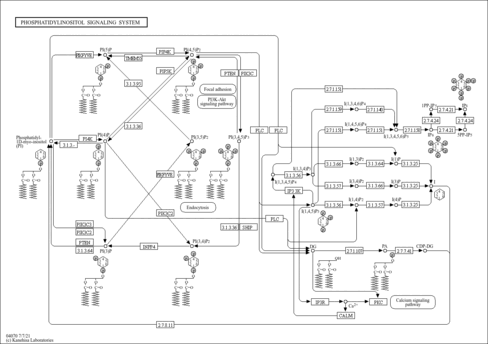 | | Inositol Phosphate Metabolism |    | 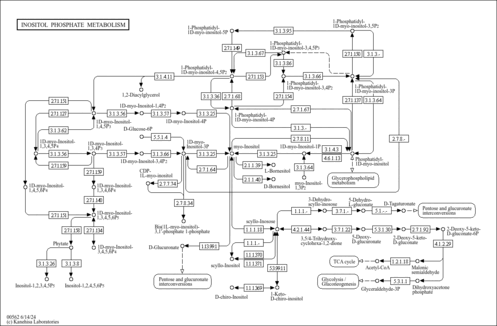 | | Inositol Metabolism |    |  |
|
| Gene Properties |
|---|
| Chromosome Location
| 8 |
| Locus
| 8q21.13-q21.3 |
| SNPs
| IMPA1 |
| Gene Sequence
|
>834 bp
ATGGCTGATCCTTGGCAGGAATGCATGGATTATGCAGTAACTCTAGCAAGACAAGCTGGA
GAGGTAGTTTGTGAAGCTATAAAAAATGAAATGAATGTTATGCTGAAAAGTTCTCCAGTT
GATTTGGTAACTGCTACGGACCAAAAAGTTGAAAAAATGCTTATCTCTTCCATAAAGGAA
AAGTATCCATCTCACAGTTTCATTGGTGAAGAATCTGTGGCAGCTGGGGAAAAAAGTATC
TTAACCGACAACCCCACATGGATCATTGACCCTATTGATGGAACAACTAACTTTGTACAT
AGATTTCCTTTTGTAGCTGTTTCAATTGGCTTTGCTGTAAATAAAAAGATAGAATTTGGA
GTTGTGTACAGTTGTGTGGAAGGCAAGATGTACACTGCCAGAAAAGGAAAAGGGGCCTTT
TGTAATGGTCAAAAACTACAAGTTTCACAACAAGAAGATATTACCAAATCTCTCTTGGTG
ACTGAGTTGGGCTCTTCTAGAACACCAGAGACTGTGAGAATGGTTCTTTCTAATATGGAA
AAGCTTTTTTGCATTCCTGTTCATGGGATCCGGAGTGTTGGAACAGCAGCTGTTAATATG
TGCCTTGTGGCAACTGGCGGAGCAGATGCATATTATGAAATGGGAATTCACTGCTGGGAT
GTTGCAGGAGCTGGCATTATTGTTACTGAAGCTGGTGGCGTGCTAATGGATGTTACAGGT
GGACCATTTGATTTGATGTCACGAAGAGTAATTGCTGCAAATAATAGAATATTAGCAGAA
AGGATAGCTAAAGAAATTCAGGTTATACCTTTGCAACGAGACGACGAAGATTAA
|
| Protein Properties |
|---|
| Number of Residues
| 277 |
| Molecular Weight
| 36694.375 |
| Theoretical pI
| 7.917 |
| Pfam Domain Function
|
|
| Signals
|
Not Available
|
|
Transmembrane Regions
|
Not Available
|
| Protein Sequence
|
>Inositol monophosphatase 1
MADPWQECMDYAVTLARQAGEVVCEAIKNEMNVMLKSSPVDLVTATDQKVEKMLISSIKE
KYPSHSFIGEESVAAGEKSILTDNPTWIIDPIDGTTNFVHRFPFVAVSIGFAVNKKIEFG
VVYSCVEGKMYTARKGKGAFCNGQKLQVSQQEDITKSLLVTELGSSRTPETVRMVLSNME
KLFCIPVHGIRSVGTAAVNMCLVATGGADAYYEMGIHCWDVAGAGIIVTEAGGVLMDVTG
GPFDLMSRRVIAANNRILAERIAKEIQVIPLQRDDED
|
| External Links |
|---|
| GenBank ID Protein
| 395340 |
| UniProtKB/Swiss-Prot ID
| P29218 |
| UniProtKB/Swiss-Prot Entry Name
| IMPA1_HUMAN |
| PDB IDs
|
|
| GenBank Gene ID
| X66922 |
| GeneCard ID
| IMPA1 |
| GenAtlas ID
| IMPA1 |
| HGNC ID
| HGNC:6050 |
| References |
|---|
| General References
| Not Available |

