| Identification |
|---|
| HMDB Protein ID
| CDBP00798 |
| Secondary Accession Numbers
| Not Available |
| Name
| Phosphoribosylformylglycinamidine synthase |
| Description
| Not Available |
| Synonyms
|
- FGAM synthase
- FGAMS
- FGARAT
- Formylglycinamide ribotide amidotransferase
- Formylglycinamide ribotide synthetase
|
| Gene Name
| PFAS |
| Protein Type
| Enzyme |
| Biological Properties |
|---|
| General Function
| Involved in catalytic activity |
| Specific Function
| Not Available |
| GO Classification
|
| Biological Process |
| purine nucleobase metabolic process |
| glutamine metabolic process |
| 'de novo' IMP biosynthetic process |
| Cellular Component |
| cytosol |
| Function |
| catalytic activity |
| ligase activity |
| carbon-nitrogen ligase activity, with glutamine as amido-n-donor |
| ligase activity, forming carbon-nitrogen bonds |
| phosphoribosylformylglycinamidine synthase activity |
| Molecular Function |
| ATP binding |
| phosphoribosylformylglycinamidine synthase activity |
| Process |
| nucleobase, nucleoside, nucleotide and nucleic acid metabolic process |
| nucleobase, nucleoside and nucleotide metabolic process |
| nucleoside phosphate metabolic process |
| nucleotide metabolic process |
| purine nucleoside monophosphate biosynthetic process |
| purine ribonucleoside monophosphate biosynthetic process |
| imp biosynthetic process |
| 'de novo' imp biosynthetic process |
| purine nucleotide metabolic process |
| purine nucleotide biosynthetic process |
| metabolic process |
| nitrogen compound metabolic process |
| cellular nitrogen compound metabolic process |
|
| Cellular Location
|
- Cytoplasm
|
| Pathways
|
| Name | SMPDB/Pathwhiz | KEGG | | IMP biosynthesis via de novo pathway | Not Available | Not Available | | Purine Metabolism |  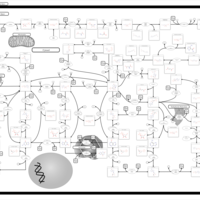 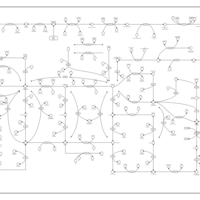 |  | | Adenosine Deaminase Deficiency |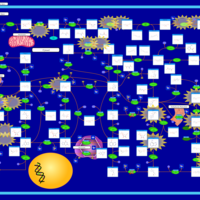  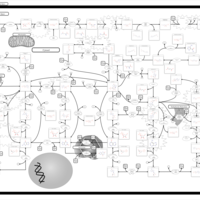 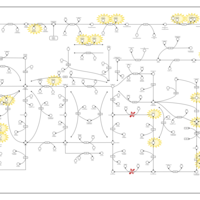 | Not Available | | Adenylosuccinate Lyase Deficiency | 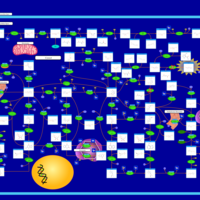 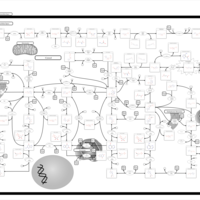 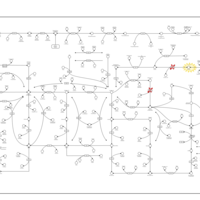 | Not Available | | Gout or Kelley-Seegmiller Syndrome | 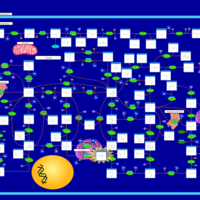 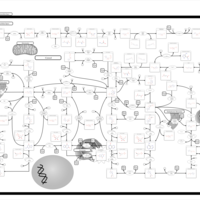 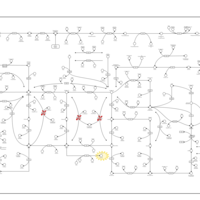 | Not Available |
|
| Gene Properties |
|---|
| Chromosome Location
| 17 |
| Locus
| 17p13.1 |
| SNPs
| PFAS |
| Gene Sequence
|
>4017 bp
ATGTCCCCAGTCCTTCACTTCTATGTTCGTCCCTCTGGCCATGAGGGGGCAGCCTCTGGA
CACACTCGGAGGAAACTGCAAGGGAAACTGCCAGAGCTGCAGGGCGTCGAGACTGAACTG
TGCTACAACGTGAACTGGACAGCTGAGGCCCTCCCCAGTGCTGAGGAGACAAAGAAGCTG
ATGTGGCTGTTTGGTTGCCCCTTACTGCTGGATGATGTTGCTCGGGAGTCCTGGCTCCTT
CCTGGCTCCAATGACCTGCTGCTGGAGGTCGGGCCCAGGCTGAACTTCTCCACCCCAACA
TCCACCAACATCGTGTCAGTGTGCCGCGCCACTGGGCTGGGGCCTGTGGATCGTGTGGAG
ACCACCCGGCGCTACCGGCTCTCGTTTGCCCACCCCCCGTCAGCTGAGGTGGAAGCCATT
GCTCTGGCTACCCTGCACGACCGGATGACAGAGCAGCACTTCCCCCATCCCATCCAGAGT
TTCTCCCCTGAGAGCATGCCGGAACCCCTCAATGGCCCTATCAATATACTGGGTGAGGGC
CGGCTTGCGCTGGAGAAGGCCAACCAGGAGCTTGGTCTGGCTTTAGACTCTTGGGACCTA
GACTTCTACACCAAGCGCTTCCAGGAGCTACAGCGGAACCCGAGCACTGTGGAGGCCTTT
GACTTGGCGCAGTCCAATAGCGAGCACAGCCGACACTGGTTCTTCAAGGGCCAGCTCCAC
GTGGATGGGCAGAAGCTGGTGCACTCACTGTTTGAGTCCATCATGAGCACCCAGGAATCC
TCGAACCCCAACAACGTCCTCAAATTCTGTGATAACAGCAGTGCAATCCAGGGAAAGGAA
GTCCGATTCCTACGGCCTGAGGACCCCACACGGCCAAGCCGCTTCCAGCAACAGCAAGGG
CTGAGACATGTTGTCTTCACAGCAGAGACTCACAACTTTCCCACAGGAGTATGCCCCTTT
AGTGGTGCAACCACTGGCACAGGGGGCCGGATTCGAGATGTCCAGTGCACAGGCCGCGGG
GCCCACGTGGTGGCTGGCACTGCCGGCTATTGCTTTGGAAATCTGCATATTCCAGGTTAC
AATCTGCCCTGGGAGGATCTAAGCTTCCAGTATCCTGGGAATTTTGCCCGGCCCCTGGAG
GTTGCCATTGAAGCCAGTAATGGAGCTTCTGACTATGGCAACAAGTTTGGGGAACCAGTG
CTGGCTGGCTTCGCCCGCTCCTTGGGCCTCCAGCTCCCAGACGGCCAGCGGCGTGAGTGG
ATCAAGCCCATCATGTTTAGTGGGGGCATTGGGTCCATGGAAGCTGACCACATAAGCAAG
GAGGCCCCAGAGCCAGGCATGGAAGTTGTAAAGGTTGGAGGTCCCGTCTACAGGATTGGA
GTTGGAGGTGGAGCTGCTTCATCTGTGCAGGTGCAGGGAGATAACACCAGTGACCTGGAC
TTTGGGGCTGTGCAGCGAGGAGACCCGGAGATGGAACAGAAGATGAACCGTGTGATCAGG
GCTTGTGTGGAGGCCCCCAAGGGAAACCCCATCTGCAGCCTTCATGATCAGGGCGCTGGT
GGCAATGGCAATGTCCTAAAAGAGCTGAGTGACCCAGCTGGAGCCATCATTTACACCAGC
CGCTTCCAGCTTGGGGACCCAACCCTGAATGCCCTGGAAATCTGGGGGGCTGAGTACCAG
GAATCAAATGCTCTTCTGCTGAGGTCCCCCAACCGGGACTTCCTGACTCATGTCAGTGCC
CGTGAACGTTGCCCGGCTTGCTTCGTGGGCACCATCACTGGAGACCGGAGAATAGTGCTG
GTGGACGATCGGGAGTGTCCTGTCAGAAGAAATGGCCAGGGGGATGCCCCCCCGACACCC
CCGCCAACCCCTGTGGACCTGGAGCTCGAATGGGTGCTGGGCAAGATGCCTCGGAAGGAG
TTCTTCCTGCAGAGGAAGCCCCCCATGCTGCAGCCTCTGGCCTTGCCCCCAGGGCTGAGC
GTGCACCAGGCTCTGGAGAGGGTTCTGAGGCTGCCCGCCGTGGCCAGCAAGCGCTACCTC
ACCAATAAGGTGGACCGCTCCGTGGGAGGCCTGGTGGCCCAGCAGCAGTGCGTGGGGCCC
CTGCAAACTCCTCTGGCAGATGTAGCGGTTGTGGCACTGAGCCATGAGGAGCTCATAGGG
GCTGCCACAGCCTTGGGAGAACAGCCAGTCAAGAGCCTGCTGGACCCAAAAGTCGCCGCC
CGGCTGGCCGTGGCCGAAGCCCTCACCAACCTGGTGTTTGCTCTGGTCACTGACCTCCGG
GATGTGAAGTGTAGCGGGAACTGGATGTGGGCAGCCAAGCTCCCAGGGGAGGGCGCAGCT
TTGGCGGATGCCTGTGAGGCTATGGTGGCAGTGATGGCAGCCCTGGGTGTGGCAGTGGAT
GGTGGCAAGGACTCCCTCAGCATGGCTGCTCGGGTTGGCACTGAGACCGTGCGGGCTCCT
GGGTCACTGGTCATCTCAGCCTATGCCGTCTGCCCAGACATCACAGCCACTGTGACCCCA
GACCTCAAGCATCCTGAAGGGAGAGGCCATCTGCTCTATGTGGCTCTGAGCCCTGGGCAG
CACCGGCTCGGGGGCACAGCTCTGGCCCAGTGCTTCTCCCAGCTTGGGGAACACCCTCCA
GACCTGGACCTTCCTGAGAACTTGGTGCGGGCCTTCAGCATCACTCAGGGGCTGCTGAAA
GACCGCCTCCTCTGCTCAGGCCACGATGTCAGTGACGGAGGCCTCGTCACATGCCTGCTG
GAGATGGCCTTTGCTGGAAATTGCGGGCTACAGGTGGATGTGCCTGTCCCCAGGGTTGAT
GTCCTGTCTGTGCTGTTCGCTGAGGAGCCAGGCCTCGTGCTGGAGGTGCAGGAGCCAGAC
CTGGCCCAGGTGCTGAAGCGTTACCGGGATGCTGGCCTCCATTGCCTGGAGCTGGGCCAC
ACAGGCGAGGCCGGGCCCCACGCCATGGTCCGGGTGTCAGTGAACGGGGCTGTGGTTCTG
GAGGAGCCTGTTGGGGAGCTGCGAGCCCTCTGGGAGGAGACGAGTTTCCAGCTGGACCGG
CTACAGGCAGAGCCTCGCTGTGTGGCAGAGGAGGAACGGGGCCTGAGGGAGCGGATGGGG
CCCAGCTATTGCCTGCCCCCCACCTTTCCCAAAGCCTCCGTGCCCCGTGAGCCTGGTGGT
CCCAGCCCCCGAGTCGCCATCTTGCGAGAGGAGGGCAGTAATGGAGACCGGGAGATGGCC
GATGCCTTCCACTTAGCTGGGTTTGAGGTATGGGACGTGACCATGCAGGACCTCTGCTCT
GGGGCAATTGGGCTGGACACTTTCCGTGGCGTGGCCTTCGTGGGCGGCTTCAGCTATGCA
GATGTCCTGGGCTCTGCCAAAGGGTGGGCAGCTGCTGTGACCTTTCATCCCAGGGCTGGG
GCTGAGCTGAGGCGCTTCCGGAAGCGGCCAGACACCTTCAGCCTGGGCGTGTGTAATGGC
TGTCAACTGCTGGCTCTGCTCGGCTGGGTGGGAGGCGACCCCAATGAGGATGCTGCAGAG
ATGGGCCCTGACTCCCAGCCAGCCCGGCCAGGCCTTCTGCTACGCCACAACCTGTCTGGG
CGCTACGAGTCTCGCTGGGCCAGCGTGCGTGTGGGGCCTGGGCCAGCCCTGATGCTGCGA
GGGATGGAGGGCGCCGTGCTGCCCGTGTGGAGTGCGCACGGGGAAGGTTACGTAGCATTT
TCTTCTCCGGAACTCCAAGCTCAGATTGAGGCCAGGGGCTTGGCTCCACTGCACTGGGCT
GATGATGACGGGAACCCCACAGAGCAGTACCCTCTGAATCCCAATGGGTCCCCAGGGGGC
GTGGCTGGCATCTGCTCCTGTGATGGCCGCCACCTGGCTGTCATGCCTCACCCTGAGCGG
GCCGTTAGGCCTTGGCAGTGGGCATGGCGACCCCCTCCATTTGATACTCTGACCACCTCC
CCCTGGCTCCAGCTCTCTATCAATGCCCGAAACTGGACCCTGGAAGGGAGCTGCTGA
|
| Protein Properties |
|---|
| Number of Residues
| 1338 |
| Molecular Weight
| 144733.165 |
| Theoretical pI
| 5.763 |
| Pfam Domain Function
|
|
| Signals
|
Not Available
|
|
Transmembrane Regions
|
Not Available
|
| Protein Sequence
|
>Phosphoribosylformylglycinamidine synthase
MSPVLHFYVRPSGHEGAASGHTRRKLQGKLPELQGVETELCYNVNWTAEALPSAEETKKL
MWLFGCPLLLDDVARESWLLPGSNDLLLEVGPRLNFSTPTSTNIVSVCRATGLGPVDRVE
TTRRYRLSFAHPPSAEVEAIALATLHDRMTEQHFPHPIQSFSPESMPEPLNGPINILGEG
RLALEKANQELGLALDSWDLDFYTKRFQELQRNPSTVEAFDLAQSNSEHSRHWFFKGQLH
VDGQKLVHSLFESIMSTQESSNPNNVLKFCDNSSAIQGKEVRFLRPEDPTRPSRFQQQQG
LRHVVFTAETHNFPTGVCPFSGATTGTGGRIRDVQCTGRGAHVVAGTAGYCFGNLHIPGY
NLPWEDPSFQYPGNFARPLEVAIEASNGASDYGNKFGEPVLAGFARSLGLQLPDGQRREW
IKPIMFSGGIGSMEADHISKEAPEPGMEVVKVGGPVYRIGVGGGAASSVQVQGDNTSDLD
FGAVQRGDPEMEQKMNRVIRACVEAPKGNPICSLHDQGAGGNGNVLKELSDPAGAIIYTS
RFQLGDPTLNALEIWGAEYQESNALLLRSPNRDFLTHVSARERCPACFVGTITGDRRIVL
VDDRECPVRRNGQGDAPPTPLPTPVDLELEWVLGKMPRKEFFLQRKPPMLQPLALPPGLS
VHQALERVLRLPAVASKRYLTNKVDRSVGGLVAQQQCVGPLQTPLADVAVVALSHEELIG
AATALGEQPVKSLLDPKVAARLAVAEALTNLVFALVTDLRDVKCSGNWMWAAKLPGEGAA
LADACEAMVAVMAALGVAVDGGKDSLSMAARVGTETVRAPGSLVISAYAVCPDITATVTP
DLKHPEGRGHLLYVALSPGQHRLGGTALAQCFSQLGEHPPDLDLPENLVRAFSITQGLLK
DRLLCSGHDVSDGGLVTCLLEMAFAGNCGLQVDVPVPRVDVLSVLFAEEPGLVLEVQEPD
LAQVLKRYRDAGLHCLELGHTGEAGPHAMVRVSVNGAVVLEEPVGELRALWEETSFQLDR
LQAEPRCVAEEERGLRERMGPSYCLPPTFPKASVPREPGGPSPRVAILREEGSNGDREMA
DAFHLAGFEVWDVTMQDLCSGAIGLDTFRGVAFVGGFSYADVLGSAKGWAAAVTFHPRAG
AELRRFRKRPDTFSLGVCNGCQLLALLGWVGGDPNEDAAEMGPDSQPARPGLLLRHNLSG
RYESRWASVRVGPGPALMLRGMEGAVLPVWSAHGEGYVAFSSPELQAQIEARGLAPLHWA
DDDGNPTEQYPLNPNGSPGGVAGICSCDGRHLAVMPHPERAVRPWQWAWRPPPFDTLTTS
PWLQLFINARNWTLEGSC
|
| External Links |
|---|
| GenBank ID Protein
| 148922280 |
| UniProtKB/Swiss-Prot ID
| O15067 |
| UniProtKB/Swiss-Prot Entry Name
| PUR4_HUMAN |
| PDB IDs
|
Not Available |
| GenBank Gene ID
| BC146768 |
| GeneCard ID
| PFAS |
| GenAtlas ID
| PFAS |
| HGNC ID
| HGNC:8863 |
| References |
|---|
| General References
| Not Available |
