| Identification |
|---|
| HMDB Protein ID
| CDBP00644 |
| Secondary Accession Numbers
| Not Available |
| Name
| Glutaminase kidney isoform, mitochondrial |
| Description
| Not Available |
| Synonyms
|
- GLS
- K-glutaminase
- L-glutamine amidohydrolase
|
| Gene Name
| GLS |
| Protein Type
| Enzyme |
| Biological Properties |
|---|
| General Function
| Involved in glutaminase activity |
| Specific Function
| Catalyzes the first reaction in the primary pathway for the renal catabolism of glutamine. Plays a role in maintaining acid-base homeostasis. Regulates the levels of the neurotransmitter glutamate in the brain. Isoform 2 lacks catalytic activity.
|
| GO Classification
|
| Biological Process |
| cellular nitrogen compound metabolic process |
| regulation of respiratory gaseous exchange by neurological system process |
| behavior |
| neurotransmitter secretion |
| protein homotetramerization |
| glutamine catabolic process |
| glutamate secretion |
| glutamate biosynthetic process |
| Cellular Component |
| cytosol |
| mitochondrial matrix |
| Function |
| hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds |
| glutaminase activity |
| hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amides |
| catalytic activity |
| hydrolase activity |
| Molecular Function |
| glutaminase activity |
| Process |
| metabolic process |
| cellular metabolic process |
| glutamine metabolic process |
| glutamine family amino acid metabolic process |
| cellular amino acid and derivative metabolic process |
| cellular amino acid metabolic process |
|
| Cellular Location
|
- Mitochondrion (Probable)
|
| Pathways
|
| Name | SMPDB/Pathwhiz | KEGG | | Alanine, aspartate and glutamate metabolism | Not Available |  | | D-Glutamine and D-glutamate metabolism | Not Available | 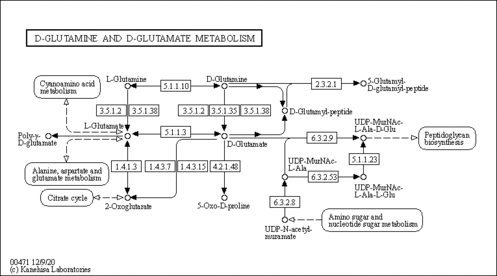 | | Glutamatergic synapse | Not Available | 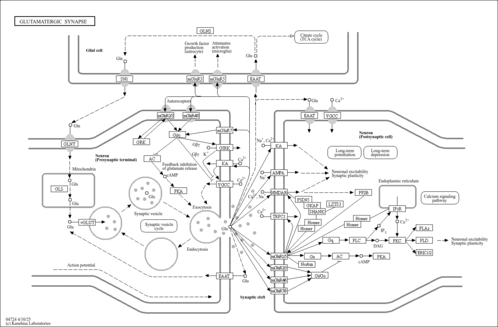 | | GABAergic synapse | Not Available | 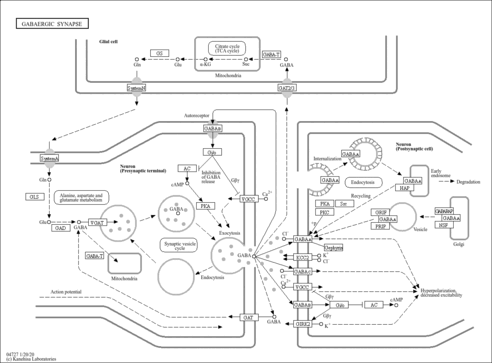 | | Proximal tubule bicarbonate reclamation | Not Available | 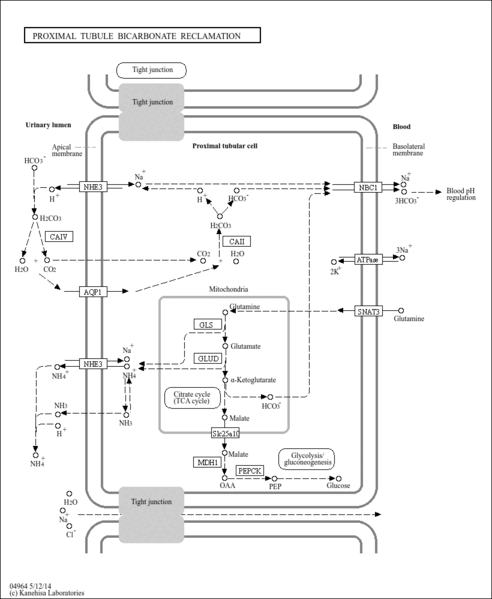 |
|
| Gene Properties |
|---|
| Chromosome Location
| 2 |
| Locus
| 2q32-q34 |
| SNPs
| GLS |
| Gene Sequence
|
>2010 bp
ATGATGCGGCTGCGAGGCTCGGGGATGCTGCGGGACCTGCTCCTGCGGTCGCCCGCCGGC
GTGAGCGCGACTCTGCGGCGGGCACAGCCCTTGGTCACCCTGTGCCGGCGTCCCCGAGGC
GGGGGACGGCCGGCCGCGGGCCCGGCTGCCGCCGCGCGACTCCACCCGTGGTGGGGCGGG
GGCGGCTGGCCGGCGGAGCCCCTCGCGCGGGGCCTGTCCAGCTCTCCTTCGGAGATCTTG
CAGGAGCTGGGCAAGGGGAGCACGCATCCGCAGCCCGGGGTGTCGCCACCCGCTGCCCCG
GCGGCGCCCGGCCCCAAGGACGGCCCCGGGGAGACGGACGCGTTTGGCAACAGCGAGGGC
AAAGAGCTGGTGGCCTCAGGTGAAAATAAAATAAAACAGGGTCTGTTACCTAGCTTGGAA
GATTTGCTGTTCTATACAATTGCTGAAGGACAAGAGAAAATACCTGTTCATAAATTTATT
ACAGCACTCAAATCTACAGGATTGCGAACGTCTGATCCCAGGTTGAAAGAGTGTATGGAT
ATGTTAAGATTAACTCTTCAAACAACATCAGATGGTGTCATGCTAGACAAAGATCTTTTT
AAAAAATGTGTTCAGAGCAACATTGTTTTGTTGACACAAGCATTTAGAAGAAAGTTTGTG
ATTCCTGACTTTATGTCTTTTACCTCACACATTGATGAGTTATATGAAAGTGCTAAAAAG
CAGTCTGGAGGAAAGGTTGCAGATTATATTCCTCAACTGGCCAAATTCAGTCCCGATTTG
TGGGGTGTGTCTGTTTGTACAGTAGATGGACAGAGGCATTCTACTGGAGATACCAAAGTT
CCCTTCTGTCTTCAGTCCTGTGTAAAACCTTTGAAATATGCCATTGCTGTTAATGATCTT
GGAACTGAATATGTGCATCGATATGTTGGAAAAGAGCCGAGTGGACTAAGATTCAACAAA
CTATTTTTGAATGAAGATGATAAACCACATAATCCTATGGTAAATGCTGGAGCAATTGTT
GTGACTTCACTAATAAAGCAAGGAGTAAATAATGCTGAAAAATTTGACTATGTCATGCAG
TTTTTGAATAAGATGGCTGGTAATGAATATGTTGGATTCAGTAATGCAACGTTTCAGTCT
GAAAGAGAAAGTGGAGATCGAAATTTTGCAATAGGATATTACTTAAAAGAAAAGAAGTGT
TTTCCAGAAGGCACAGACATGGTTGGTATATTAGACTTCTACTTCCAGCTGTGCTCCATT
GAAGTGACTTGTGAATCAGCCAGTGTGATGGCTGCGACACTGGCTAATGGTGGTTTCTGC
CCAATTACTGGTGAAAGAGTACTGAGCCCTGAAGCAGTTCGAAATACATTGAGTTTGATG
CATTCCTGTGGCATGTATGACTTCTCAGGGCAGTTTGCTTTCCATGTTGGTCTTCCTGCA
AAATCTGGAGTTGCTGGGGGCATTCTTTTAGTTGTCCCCAATGTTATGGGTATGATGTGC
TGGTCTCCTCCTCTGGATAAGATGGGCAACAGTGTTAAGGGAATTCACTTTTGTCACGAT
CTTGTTTCTCTGTGTAATTTCCATAACTATGATAATTTGAGACACTTTGCAAAAAAACTT
GATCCTCGAAGAGAAGGTGGTGATCAAAGGGTAAAGTCAGTGATAAATCTTTTGTTTGCT
GCATATACTGGAGATGTGTCTGCACTTCGAAGATTTGCTTTGTCAGCTATGGACATGGAA
CAGCGGGACTATGATTCTAGAACAGCACTCCATGTAGCTGCTGCAGAGGGTCATGTTGAA
GTTGTTAAATTTTTGCTGGAAGCCTGCAAAGTAAACCCTTTCCCCAAGGACAGGTGGAAT
AACACTCCCATGGATGAAGCACTGCACTTTGGACACCATGATGTATTTAAAATTCTCCAA
GAATACCAAGTCCAGTACACACCTCAAGGAGATTCTGACAACGGGAAGGAAAATCAAACC
GTCCATAAGAATCTTGATGGATTGTTGTAA
|
| Protein Properties |
|---|
| Number of Residues
| 669 |
| Molecular Weight
| 65459.525 |
| Theoretical pI
| 7.885 |
| Pfam Domain Function
|
|
| Signals
|
Not Available
|
|
Transmembrane Regions
|
Not Available
|
| Protein Sequence
|
>Glutaminase kidney isoform, mitochondrial
MMRLRGSGMLRDLLLRSPAGVSATLRRAQPLVTLCRRPRGGGRPAAGPAAAARLHPWWGG
GGWPAEPLARGLSSSPSEILQELGKGSTHPQPGVSPPAAPAAPGPKDGPGETDAFGNSEG
KELVASGENKIKQGLLPSLEDLLFYTIAEGQEKIPVHKFITALKSTGLRTSDPRLKECMD
MLRLTLQTTSDGVMLDKDLFKKCVQSNIVLLTQAFRRKFVIPDFMSFTSHIDELYESAKK
QSGGKVADYIPQLAKFSPDLWGVSVCTVDGQRHSTGDTKVPFCLQSCVKPLKYAIAVNDL
GTEYVHRYVGKEPSGLRFNKLFLNEDDKPHNPMVNAGAIVVTSLIKQGVNNAEKFDYVMQ
FLNKMAGNEYVGFSNATFQSERESGDRNFAIGYYLKEKKCFPEGTDMVGILDFYFQLCSI
EVTCESASVMAATLANGGFCPITGERVLSPEAVRNTLSLMHSCGMYDFSGQFAFHVGLPA
KSGVAGGILLVVPNVMGMMCWSPPLDKMGNSVKGIHFCHDLVSLCNFHNYDNLRHFAKKL
DPRREGGDQRVKSVINLLFAAYTGDVSALRRFALSAMDMEQRDYDSRTALHVAAAEGHVE
VVKFLLEACKVNPFPKDRWNNTPMDEALHFGHHDVFKILQEYQVQYTPQGDSDNGKENQT
VHKNLDGLL
|
| External Links |
|---|
| GenBank ID Protein
| 156104878 |
| UniProtKB/Swiss-Prot ID
| O94925 |
| UniProtKB/Swiss-Prot Entry Name
| GLSK_HUMAN |
| PDB IDs
|
|
| GenBank Gene ID
| NM_014905.3 |
| GeneCard ID
| GLS |
| GenAtlas ID
| GLS |
| HGNC ID
| HGNC:4331 |
| References |
|---|
| General References
| Not Available |




