| Record Information |
|---|
| Version | 1.0 |
|---|
| Created at | 2020-04-17 19:21:53 UTC |
|---|
| Updated at | 2021-01-04 18:49:00 UTC |
|---|
| CannabisDB ID | CDB005225 |
|---|
| Secondary Accession Numbers | Not Available |
|---|
| Cannabis Compound Identification |
|---|
| Common Name | Squalene |
|---|
| Description | Squalene is an unsaturated aliphatic hydrocarbon (carotenoid) with six unconjugated double bonds. It belongs to the class of compounds known as triterpenoids. These are a class of chemical compounds composed of three terpene units with the molecular formula C30H48; they may also be thought of as consisting of six isoprene units. Triterpenes are biosynthesized through the head-to-head condensation of two FPP units to form squalene. In turn, squalene serves as precursor for the formation of triterpenoids, including bacterial hopanoids and eukaryotic sterols. Squalene is a natural organic compound originally obtained for commercial purposes primarily from shark liver oil (hence its name, as Squalus is a genus of sharks). Squalene is a low-density compound that is stored in the bodies of cartilaginous fishes such as sharks, which lack a swim bladder and must therefore reduce their body density with fats and oils. Squalene, which is stored mainly in the shark's liver, is lighter than water with a specific gravity of 0.855 (Wikipedia ). Squalene can also be isolated through botanical sources as well, including rice bran, wheat germ, and olives. It occurs in high concentrations in the stomach oil of birds in the order Procellariiformes.Industrially, squalene is used as a bactericide. It is also an intermediate in the manufacture of pharmaceuticals, rubber chemicals, and colouring materials. All higher organisms produce squalene, including yeast, plants, insects, fish and humans. However, blue-green algae and some bacteria do not manufacture squalene and must acquire it from the environment if they need it. Squalene may function as a pheromone and is a volatile component of the scent material from Saguinus oedipus (cotton-top tamarin monkey) and Saguinus fuscicollis (saddle-back tamarin monkey) It is also found in human sebum (5%), fish liver oils, yeast lipids, and many vegetable oils (e.g. palm oil, cottonseed oil, rapeseed oil). Squalene is a biochemical precursor to nearly all steroids. Oxidation of one of the terminal double bonds of squalene yields 2,3-squalene oxide which undergoes enzyme-catalyzed cyclization to afford lanosterol, which is then elaborated into cholesterol and other steroids. Squalene is also a constituent of cannabis smoke and is formed during the combustion of cannabis ( Ref:DOI ). |
|---|
| Structure | |
|---|
| Synonyms | | Value | Source |
|---|
| (all-e)-2,6,10,15,19,23-Hexamethyl-2,6,10,14,18,22-tetracosahexaene | ChEBI | | Spinacene | ChEBI | | Supraene | ChEBI | | (e,e,e,e)-Squalene | HMDB | | all-trans-Squalene | HMDB | | trans-Squalene | HMDB | | Squalene | HMDB |
|
|---|
| Chemical Formula | C30H50 |
|---|
| Average Molecular Weight | 410.73 |
|---|
| Monoisotopic Molecular Weight | 410.3913 |
|---|
| IUPAC Name | (6E,10E,14E,18E)-2,6,10,15,19,23-hexamethyltetracosa-2,6,10,14,18,22-hexaene |
|---|
| Traditional Name | squalene |
|---|
| CAS Registry Number | 111-02-4 |
|---|
| SMILES | CC(C)=CCC\C(C)=C\CC\C(C)=C\CC\C=C(/C)CC\C=C(/C)CCC=C(C)C |
|---|
| InChI Identifier | InChI=1S/C30H50/c1-25(2)15-11-19-29(7)23-13-21-27(5)17-9-10-18-28(6)22-14-24-30(8)20-12-16-26(3)4/h15-18,23-24H,9-14,19-22H2,1-8H3/b27-17+,28-18+,29-23+,30-24+ |
|---|
| InChI Key | YYGNTYWPHWGJRM-AAJYLUCBSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | Belongs to the class of organic compounds known as triterpenoids. These are terpene molecules containing six isoprene units. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Prenol lipids |
|---|
| Sub Class | Triterpenoids |
|---|
| Direct Parent | Triterpenoids |
|---|
| Alternative Parents | |
|---|
| Substituents | - Triterpenoid
- Branched unsaturated hydrocarbon
- Unsaturated aliphatic hydrocarbon
- Unsaturated hydrocarbon
- Olefin
- Acyclic olefin
- Hydrocarbon
- Aliphatic acyclic compound
|
|---|
| Molecular Framework | Aliphatic acyclic compounds |
|---|
| External Descriptors | |
|---|
| Ontology |
|---|
|
| Disposition | Route of exposure: Source: Biological location: |
|---|
| Role | Industrial application: Biological role: |
|---|
| Physical Properties |
|---|
| State | Liquid |
|---|
| Experimental Properties | | Property | Value | Reference |
|---|
| Melting Point | -75 °C | Not Available | | Boiling Point | 285 °C | Wikipedia | | Water Solubility | Not Available | Not Available | | logP | 12.188 | Wikipedia |
|
|---|
| Predicted Properties | [] |
|---|
| Spectra |
|---|
| EI-MS/GC-MS | | Type | Description | Splash Key | View |
|---|
| EI-MS | Mass Spectrum (Electron Ionization) | splash10-0159-9610000000-aaf0145da95343082ae1 | 2015-03-01 | View Spectrum | | GC-MS | Squalene, non-derivatized, GC-MS Spectrum | splash10-001i-9600000000-280b7ccaed0215f0937c | Spectrum | | GC-MS | Squalene, non-derivatized, GC-MS Spectrum | splash10-015i-9500000000-63fc959bf17e2f6e0381 | Spectrum | | Predicted GC-MS | Squalene, non-derivatized, Predicted GC-MS Spectrum - 70eV, Positive | splash10-05dv-4879000000-3cfdaa630f73c021be06 | Spectrum | | Predicted GC-MS | Squalene, non-derivatized, Predicted GC-MS Spectrum - 70eV, Positive | Not Available | Spectrum |
|
|---|
| MS/MS | | Type | Description | Splash Key | View |
|---|
| MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 10V, Positive (Annotated) | splash10-047r-1944600000-5faa7a5eb4804dfad261 | 2019-11-13 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 25V, Positive (Annotated) | splash10-000i-4867900000-dcd3ba4f6ee550dc565b | 2019-11-13 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - Quattro_QQQ 40V, Positive (Annotated) | splash10-000i-2944400000-8be7743f6b07ba138c24 | 2019-11-13 | View Spectrum | | MS/MS | LC-MS/MS Spectrum - 20V, Positive | splash10-03dj-2920300000-04f33a28f9d38283345e | 2021-09-20 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-03di-0323900000-9162860711ec9f8266fe | 2016-08-03 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-02fx-2984100000-742302f14e45ce03bfdd | 2016-08-03 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-014l-6495000000-83c6df474790679c40e3 | 2016-08-03 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0a4i-0000900000-4905353fab78414bcab2 | 2016-08-03 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0a4i-0000900000-e7195906634d18bf31dd | 2016-08-03 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-0006-1869100000-2d7d4eb74fe0c5856cbf | 2016-08-03 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 10V, Positive | splash10-03di-2844900000-f00046ef895fbe9888a3 | 2021-09-24 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 20V, Positive | splash10-05mk-4928000000-3dfbc6d8fc1271828c3b | 2021-09-24 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 40V, Positive | splash10-0a5j-4932000000-6f14fcc4a6e8029a1652 | 2021-09-24 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 10V, Negative | splash10-0a4i-0000900000-45d1d2cb6b79e3bc92ae | 2021-09-24 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 20V, Negative | splash10-0a4i-0102900000-49aa25403f6062cc3520 | 2021-09-24 | View Spectrum | | Predicted MS/MS | Predicted LC-MS/MS Spectrum - 40V, Negative | splash10-002o-1239000000-809fbf3fc0f0117fe5d1 | 2021-09-24 | View Spectrum |
|
|---|
| NMR | | Type | Description | | View |
|---|
| 1D NMR | 13C NMR Spectrum (1D, 25.16 MHz, CDCl3, experimental) | | Spectrum | | 2D NMR | [1H, 13C]-HSQC NMR Spectrum (2D, 600 MHz, CDCl3, experimental) | | Spectrum |
|
|---|
| Pathways |
|---|
| Pathways | | Name | SMPDB/Pathwhiz | KEGG | | Steroid Biosynthesis | 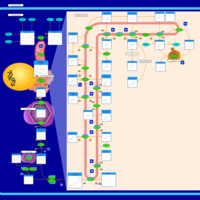 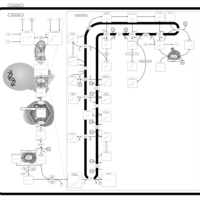 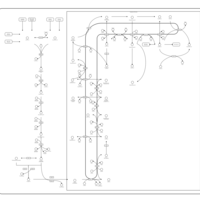 | 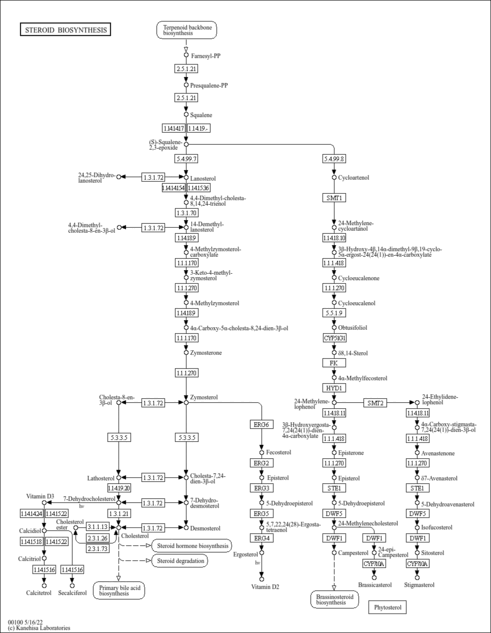 | | Smith-Lemli-Opitz Syndrome (SLOS) | 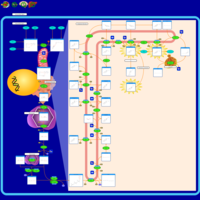 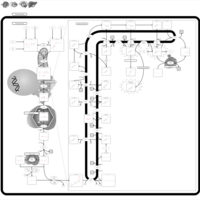 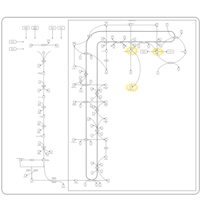 | Not Available | | CHILD Syndrome | 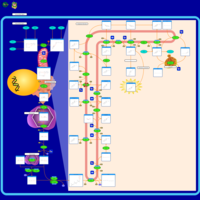 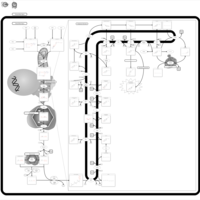 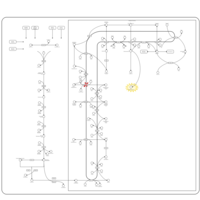 | Not Available | | Desmosterolosis | 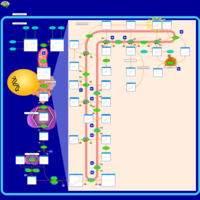 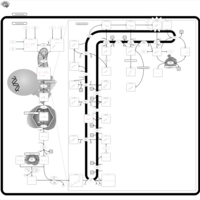 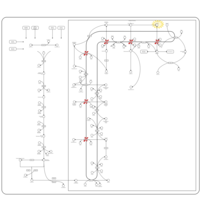 | Not Available | | Chondrodysplasia Punctata II, X Linked Dominant (CDPX2) | 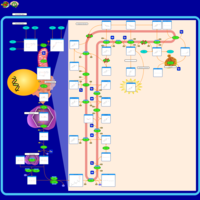 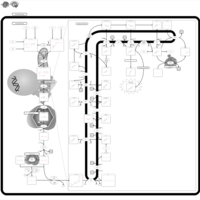 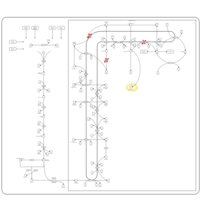 | Not Available |
|
|---|
| Protein Targets |
|---|
| Enzymes | |
|---|
| Transporters | |
|---|
| Metal Bindings | Not Available |
|---|
| Receptors | Not Available |
|---|
| Transcriptional Factors | Not Available |
|---|
| Concentrations Data |
|---|
| Not Available |
|---|
| External Links |
|---|
| HMDB ID | HMDB0000256 |
|---|
| DrugBank ID | DB11460 |
|---|
| Phenol Explorer Compound ID | Not Available |
|---|
| FoodDB ID | Not Available |
|---|
| KNApSAcK ID | C00003755 |
|---|
| Chemspider ID | 553635 |
|---|
| KEGG Compound ID | C00751 |
|---|
| BioCyc ID | SQUALENE |
|---|
| BiGG ID | Not Available |
|---|
| Wikipedia Link | Squalene |
|---|
| METLIN ID | Not Available |
|---|
| PubChem Compound | 638072 |
|---|
| PDB ID | Not Available |
|---|
| ChEBI ID | 15440 |
|---|
| References |
|---|
| General References | Not Available |
|---|
